Overview
Source: Laboratory of Jeff Salacup - University of Massachusetts Amherst
As mentioned in previous videos, the product of an organic solvent extraction, a total lipid extract (TLE), is often a complex mixture of hundreds, if not thousands, of different compounds. The researcher is often only interested in a handful of compounds. In the case of our two organic paleothermometers (Uk'37 and MBT/CBT), the interest is in only 6 compounds (2 alkenones and 4 isoprenoidal glycerol dialkyl glycerol tetraethers). As discussed in the previous two videos in this series, purification techniques may be applied in order to pare down the number of compounds in an analyzed sample. These techniques may chemically alter the unwanted components (saponification), take advantage of the different compound chemistries (column chromatography), or use the different shapes and sizes of the molecules to include or exclude certain components from the analysis (urea adduction). The atomic structure of different chemicals leads some organic compounds to form long, narrow, straight chains (n-alkanes and alkenones), other organic compounds to form complex cyclic structures, others to form highly-branched structures, and yet others which form both cyclic and branched structures (GDGTs) (Figure 1). The different shapes and sizes of the compounds in a sample can be used to separate them from one another, in much the same way as a coin sorter separates coins of different denominations (sizes).
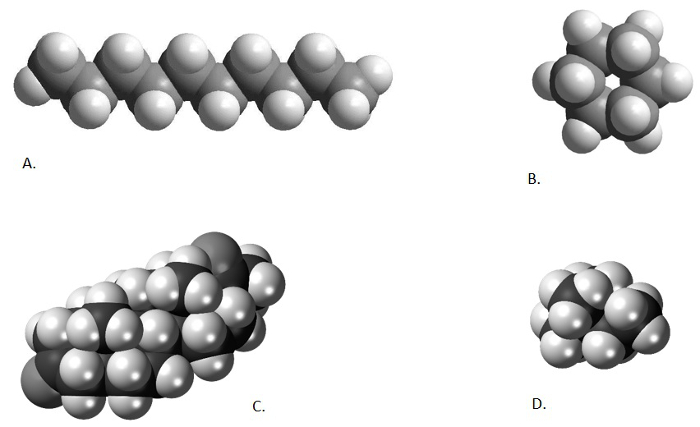
Figure 1. Comparison of different chemical structures. Decane, a straight-chained alkane (A; from from http://www.bpc.edu/mathscience/chemistry), cyclohexane, a cyclic alkane (B; from http://www.bpc.edu/mathscience/chemistry), a steroid, a poly-cyclic hydrocarbon (C; from www.wikiwand.com), and 2,2-dimethylbutane, a branched alkane (D; www.wikimedia.com). Please click here to view a larger version of this figure.
Principles
Urea adduction is a size exclusion technique that separates straight-chained or rarely-branching structures from highly-branched and cyclic structures. This happens because of the special crystal structure of urea (Figure 2). When a urea crystal forms, it creates tiny spaces between the individual crystals. The spaces are long and narrow with an average diameter of five angstroms (5 x 10-10 m). These spaces are large enough to include straight-chain or rarely-branching compounds into the crystal lattice, but too small for highly-branched or cyclic structures. Thus, the latter structures are excluded. The crystals can then be washed, and the highly-branched or cyclic structures can be removed from the sample. Subsequent dissolution of the crystal releases the straight-chained and rarely-branching compounds back into the solution, from which they can be extracted and analyzed.

Figure 2. Depiction of the chemical and crystal structure of urea. (from www.imgkid.com) Please click here to view a larger version of this figure.
Urea adduction is a technique that separates straight-chained alkenones from highly-branched and cyclic compounds.
Urea is an organic compound that forms a porous crystalline structure. The crystal can trap certain molecules, forming an adduct, while others can’t fit within the structure.
Urea adduction utilizes the different sizes and shapes of compounds to separate them – similar to how a coin counting machine will sort a jar of loose change.
This video is part of a series on lipid extraction, purification, and analysis from sediments. It will illustrate the use of urea adduction to perform size exclusion purification on a sample for alkenone paleothermometry.
Urea adduction is a purification method based on size exclusion. It separates straight-chained or rarely-branching compounds – such as n-alkanes and alkenones - from those that are highly-branched and/or cyclic.
This is possible because of urea's special crystalline structure. When a urea crystal forms, tiny spaces are created between the individual molecules. These spaces are long and narrow - so straight-chain or rarely-branching compounds can fit into the crystal lattice, while highly-branched and cyclic compounds are too large.
The urea crystals are then washed with an apolar solvent, separating the excluded molecules from the included linear ones. The washed molecules can be extracted and analyzed directly, while the crystals must be dissolved in water to release the linear compounds back into solution first. Another apolar solvent is then used to extract the desired compounds from the water.
Both the included and excluded molecules can provide valuable information. For example, highly branched isoprenoids, produced by sea ice diatoms, can be a proxy for the existence of seasonal sea ice at high latitudes. Cyclic compounds may reflect the presence of past fires. Straight chained alkanes and alkenones are common proxies for ecosystem structure and sea surface temperature.
The purification provided by urea adduction is not always necessary, as some compounds of interest can be analyzed directly from the unaltered extracted organic sample. In extreme cases - such as in sediments acquired from highly polluted areas, like estuaries near industrial centers - a urea adduction may be necessary to remove unknown compounds that coelute during analysis.
Now that you understand urea adduction, you are ready to begin the procedure.
To begin, acquire a dried total lipid extract – or TLE – that has been extracted from the sample and purified with saponification and column chromatography.
Next, prepare the urea adduction solutions as outlined in the text protocol. Ensure that all components are pure and free of hydrocarbons.
Suspend the sample in 1.5 mL of the DCM/hexane solution. If the TLE does not completely dissolve, sonicate for 5 min. Add 1.5 mL of the urea and methanol solution. Watch for the formation of a white precipitate, as this signals the creation of urea crystals. Next, gently dry the urea crystals under nitrogen, using gentle heat. Be sure to evaporate all of the solvent. Once the crystals are completely dried, rinse them 3 times with approximately 1 mL of hexane. Use a glass pipette to remove the hexane between each rinse, and transfer it to a fresh vial. This is labeled the "non-adduct". Next, dissolve the crystals in 2 mL of pure water. Shake to ensure complete dissolution.
To extract the biomarkers from the added water, add 1 mL of hexane, cap, and gently shake for 5 s.
Allow the solution to rest until the hexane and water separate completely. Then, using a glass pipette, remove approximately 75% of the hexane and transfer it into a new vial. This is labeled the "adduct".
Repeat this extraction process twice, adding 1 mL of hexane each time. Combine the three adducts into one vial. Dispose of the water and urea solution in an appropriate waste container. The sample is now ready for analysis.
Urea adduction has several applications in the separation and purification of organic molecules.
An isotope is a variant of a chemical element that differs in the neutron quantity, and thus differs in its atomic mass. An element may have several isotopes, each having a different mass. Isotopes can also have chemical and molecular properties that differ from one another, so it can be important to determine which isotopes are present in a particular sample. In this example, the carbon and hydrogen isotope ratios in leaf waxes were measured in order to gather information on a plant's metabolic pathways. Very large quantities of material are required to determine isotope ratios, and compounds that have similar detections at low concentrations may overlap when large quantities are analyzed.
Therefore, urea adduction was used to separate the n-alkanes of interest from any interfering compounds. Removing these unwanted materials allowed for an accurate isotope ratio to be determined.
Petroleum is a complex mixture of hydrocarbons, each with unique properties and uses. The separation of these compounds from petroleum is very important in the chemical industry - branched-chain alkanes are often used as light lubricants, while straight-chain alkanes are mainly used in alkylation processes. In this example, urea adduction was used to separate alkanes from kerosene. Using a successive series of urea adductions, these alkanes were separated from kerosene at a 99-percentage purity.
You’ve just watched JoVE’s introduction to the purification of complex organic mixtures via urea adduction. You should now understand size exclusion, the importance of purifying samples for accurate component measurement, and urea adduction.
Thanks for watching!
Subscription Required. Please recommend JoVE to your librarian.
Procedure
1. Setup and Preparation of Materials
- Obtain a total lipid extract (TLE) using a solvent extraction method (Sonication, Soxhlet, or Accelerated Solvent Extraction (ASE)).
- Purchase the following materials from any chemical retailer: combusted borosilicate glass pipettes and bulbs; pure water; hexane; Dichloromethane (DCM); methanol; urea.
- The reagents should be pure and free from hydrocarbons. Alternatively, pure water can be made in a lab using a water purification system.
- Obtain 4-mL borosilicate glass vials with PTFE-lined caps.
2. Methods
- Make a mixture of 2:1 DCM:Hexane (v:v).
- In an Erlenmeyer flask, mix 200 mL of DCM and 100 mL of hexane.
- Make a mixture of urea in methanol (100 mg/mL).
- Pour 300 mL of methanol into an Erlenmeyer flask.
- Add 300 mg of urea.
- Stir on an automatic stir plate until the urea is completely dissolved.
- Start with the dry sample in a 4-mL vial.
- Suspend the sample in 1.5 mL of a 2:1 mixture of DCM:Hexane (v:v).
- If the sample does not completely dissolve, sonicate for 5 min.
- To this mixture, add 1.5 mL of urea in methanol.
- After this addition, watch for a white precipitate to form. This signals the creation of urea crystals.
- Gently dry the crystals under nitrogen, using gentle heat. Ensure all solvent has evaporated before proceeding to the next step.
- Rinse the crystals 3x with approximately 1 mL of hexane. Remove the hexane between each rinse using a glass pipette to transfer it into a vial labeled "non-adduct."
- Dissolve the crystals in 2 mL of pure water. Shake to ensure complete dissolution.
- Extract the aqueous phase with hexane by adding 1 mL and gently shaking for 5 s.
- Allow the hexane and water to completely separate before removing approximately 75% of the hexane using a glass pipette and transferring it into a vial labeled "adduct."
- Repeat step 2.10 twice.
- Dispose of the water and urea solution in an appropriate waste container.
Urea adduction is a technique that separates straight-chained alkenones from highly-branched and cyclic compounds.
Urea is an organic compound that forms a porous crystalline structure. The crystal can trap certain molecules, forming an adduct, while others can’t fit within the structure.
Urea adduction utilizes the different sizes and shapes of compounds to separate them – similar to how a coin counting machine will sort a jar of loose change.
This video is part of a series on lipid extraction, purification, and analysis from sediments. It will illustrate the use of urea adduction to perform size exclusion purification on a sample for alkenone paleothermometry.
Urea adduction is a purification method based on size exclusion. It separates straight-chained or rarely-branching compounds – such as n-alkanes and alkenones - from those that are highly-branched and/or cyclic.
This is possible because of urea's special crystalline structure. When a urea crystal forms, tiny spaces are created between the individual molecules. These spaces are long and narrow - so straight-chain or rarely-branching compounds can fit into the crystal lattice, while highly-branched and cyclic compounds are too large.
The urea crystals are then washed with an apolar solvent, separating the excluded molecules from the included linear ones. The washed molecules can be extracted and analyzed directly, while the crystals must be dissolved in water to release the linear compounds back into solution first. Another apolar solvent is then used to extract the desired compounds from the water.
Both the included and excluded molecules can provide valuable information. For example, highly branched isoprenoids, produced by sea ice diatoms, can be a proxy for the existence of seasonal sea ice at high latitudes. Cyclic compounds may reflect the presence of past fires. Straight chained alkanes and alkenones are common proxies for ecosystem structure and sea surface temperature.
The purification provided by urea adduction is not always necessary, as some compounds of interest can be analyzed directly from the unaltered extracted organic sample. In extreme cases - such as in sediments acquired from highly polluted areas, like estuaries near industrial centers - a urea adduction may be necessary to remove unknown compounds that coelute during analysis.
Now that you understand urea adduction, you are ready to begin the procedure.
To begin, acquire a dried total lipid extract – or TLE – that has been extracted from the sample and purified with saponification and column chromatography.
Next, prepare the urea adduction solutions as outlined in the text protocol. Ensure that all components are pure and free of hydrocarbons.
Suspend the sample in 1.5 mL of the DCM/hexane solution. If the TLE does not completely dissolve, sonicate for 5 min. Add 1.5 mL of the urea and methanol solution. Watch for the formation of a white precipitate, as this signals the creation of urea crystals. Next, gently dry the urea crystals under nitrogen, using gentle heat. Be sure to evaporate all of the solvent. Once the crystals are completely dried, rinse them 3 times with approximately 1 mL of hexane. Use a glass pipette to remove the hexane between each rinse, and transfer it to a fresh vial. This is labeled the "non-adduct". Next, dissolve the crystals in 2 mL of pure water. Shake to ensure complete dissolution.
To extract the biomarkers from the added water, add 1 mL of hexane, cap, and gently shake for 5 s.
Allow the solution to rest until the hexane and water separate completely. Then, using a glass pipette, remove approximately 75% of the hexane and transfer it into a new vial. This is labeled the "adduct".
Repeat this extraction process twice, adding 1 mL of hexane each time. Combine the three adducts into one vial. Dispose of the water and urea solution in an appropriate waste container. The sample is now ready for analysis.
Urea adduction has several applications in the separation and purification of organic molecules.
An isotope is a variant of a chemical element that differs in the neutron quantity, and thus differs in its atomic mass. An element may have several isotopes, each having a different mass. Isotopes can also have chemical and molecular properties that differ from one another, so it can be important to determine which isotopes are present in a particular sample. In this example, the carbon and hydrogen isotope ratios in leaf waxes were measured in order to gather information on a plant's metabolic pathways. Very large quantities of material are required to determine isotope ratios, and compounds that have similar detections at low concentrations may overlap when large quantities are analyzed.
Therefore, urea adduction was used to separate the n-alkanes of interest from any interfering compounds. Removing these unwanted materials allowed for an accurate isotope ratio to be determined.
Petroleum is a complex mixture of hydrocarbons, each with unique properties and uses. The separation of these compounds from petroleum is very important in the chemical industry - branched-chain alkanes are often used as light lubricants, while straight-chain alkanes are mainly used in alkylation processes. In this example, urea adduction was used to separate alkanes from kerosene. Using a successive series of urea adductions, these alkanes were separated from kerosene at a 99-percentage purity.
You’ve just watched JoVE’s introduction to the purification of complex organic mixtures via urea adduction. You should now understand size exclusion, the importance of purifying samples for accurate component measurement, and urea adduction.
Thanks for watching!
Subscription Required. Please recommend JoVE to your librarian.
Results
This purification technique produces two different vials; one labeled adduct, containing straight-chained and rarely-branching compounds, and another labeled non-adduct, containing highly-branched and cyclic compounds. This procedure has vastly decreased the complexity of any sample to be analyzed on an instrument. This decrease in complexity is often crucial to the accurate analysis of target compounds. For example, in nearshore settings after approximately 1850, alkenones co-elute with troublesome compounds, ostensibly pollutants introduced after the Industrial Revolution, which are not removed using column chromatography or saponification alone. Apparently, the pollutants are cyclic or branched because urea adduction removes them from the sample. The top 160 years of some of these sediment archives can be confidently analyzed for Uk'37 only because of the application of urea adduction.
Subscription Required. Please recommend JoVE to your librarian.
Applications and Summary
Urea adduction is often used in the purification of n-alkanes, common constituents of leaf wax, in order to remove co-eluting compounds before isotope analysis. The carbon and hydrogen isotope ratios of leaf waxes in plants contain information on the metabolic pathways and environmental conditions the plant used and lived in, respectively. In order to determine the isotope ratios, very large quantities of compound must be loaded onto a GC. Such large quantities often cause compounds that elute close to one another at lower concentrations to co-elute. Often, the compounds co-eluting with alkanes are branched or cyclic moieties of that alkane and can thus be removed using urea adduction.
Subscription Required. Please recommend JoVE to your librarian.




