
Proteomics Analyses Using OpenProt Database
Xavier Roucou’s lab, at the Université de Sherbrooke, presents a protocol for using OpenProt databases when interrogating mass spectrometry datasets.
Comparing Protein Levels with Western Blotting

Thomas H. Gillingwater’s lab, at the University of Edinburgh, describes an approach for comparing protein levels across tissues and through development using standardized quantitative western blotting.
Working With Drosophila Cell Lines

Arthur Luhur and colleagues, at Indiana University Bloomington, demonstrate protocols for thawing, subculturing, and cryopreserving Drosophila cell lines.
Manipulating Gene Function in Cavefish

Erik R. Duboué’s lab, at Florida Atlantic University, presents approaches for the manipulation of genes in the evolutionary model system Astyanax mexicanus.
Quantifying mRNAs With SM-FISH
Jennifer R. Wood’s lab, at the University of Nebraska-Lincoln, demonstrates a protocol for counting the number of mRNAs in individual murine oocytes using single molecule RNA fluorescence in situ hybridization.
Bile Duct Density Determination

Hamed Jafar-Nejad’s lab, at Baylor College of Medicine, describes a method for quantifying the bile duct density in the mouse liver for determining effects of genetic and environmental modifiers.

Open for Submissions Now
Current Methods for Membrane Receptor Probing in the Vascular System
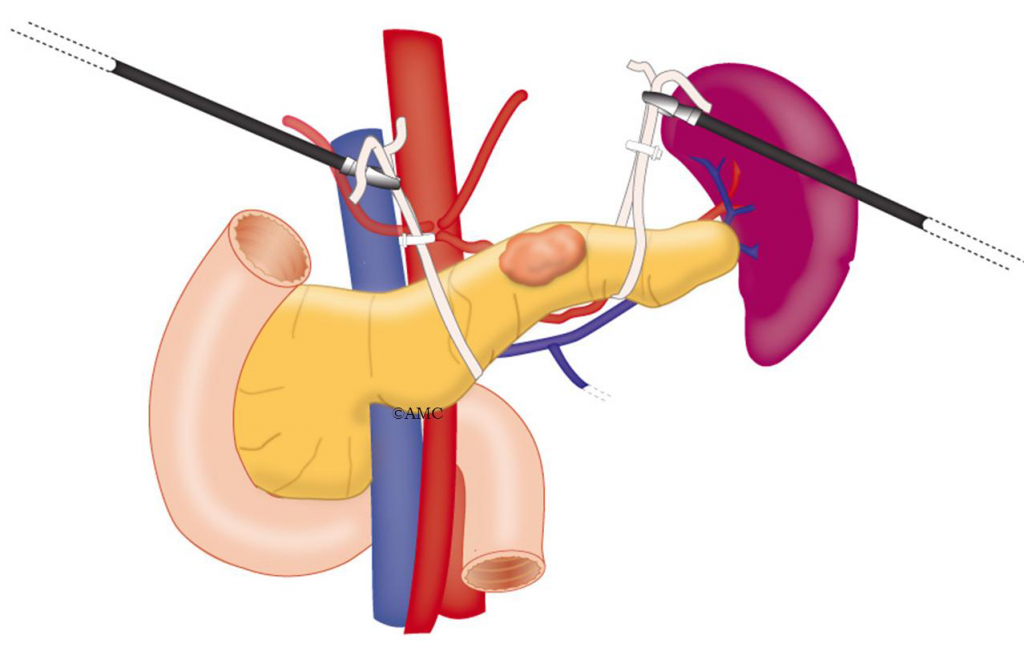
Minimally Invasive Pancreatic Surgery


This collection focuses on genetics and incorporates five broad subdisciplines: the genetics of individuals and populations; genetics and disease; gene expression; epigenetics; and genetic engineering.
Biology I: Yeast, Drosophila, and C. elegans

This collection features three model organisms commonly used in life sciences research, and also covers methodologies to maintain them in the laboratory.
This collection introduces the field of developmental biology and covers five areas, including developmental genetics and stem cell biology.

This collection provides both general and specific safety guidelines to be followed when working with hazardous materials and equipment.





