Electrophoretic Mobility Shift Assay (EMSA) for the Study of RNA-Protein Interactions: The IRE/IRP Example
Summary
Here we present a protocol to analyze RNA/protein interactions. The electrophoretic mobility shift assay (EMSA) is based on the differential migration of RNA/protein complexes and free RNA during native gel electrophoresis. By using a radiolabeled RNA probe, RNA/protein complexes can be visualized by autoradiography.
Abstract
RNA/protein interactions are critical for post-transcriptional regulatory pathways. Among the best-characterized cytosolic RNA-binding proteins are iron regulatory proteins, IRP1 and IRP2. They bind to iron responsive elements (IREs) within the untranslated regions (UTRs) of several target mRNAs, thereby controlling the mRNAs translation or stability. IRE/IRP interactions have been widely studied by EMSA. Here, we describe the EMSA protocol for analyzing the IRE-binding activity of IRP1 and IRP2, which can be generalized to assess the activity of other RNA-binding proteins as well. A crude protein lysate containing an RNA-binding protein, or a purified preparation of this protein, is incubated with an excess of32 P-labeled RNA probe, allowing for complex formation. Heparin is added to preclude non-specific protein to probe binding. Subsequently, the mixture is analyzed by non-denaturing electrophoresis on a polyacrylamide gel. The free probe migrates fast, while the RNA/protein complex exhibits retarded mobility; hence, the procedure is also called “gel retardation” or “bandshift” assay. After completion of the electrophoresis, the gel is dried and RNA/protein complexes, as well as free probe, are detected by autoradiography. The overall goal of the protocol is to detect and quantify IRE/IRP and other RNA/protein interactions. Moreover, EMSA can also be used to determine specificity, binding affinity, and stoichiometry of the RNA/protein interaction under investigation.
Introduction
The EMSA was originally developed to study the association of DNA-binding proteins with target DNA sequences1,2. The principle is similar for RNA/protein interactions3, which is the focus of this article. Briefly, RNA is negatively charged and will migrate towards the anode during non-denaturing electrophoresis in polyacrylamide (or agarose) gels. Migration within the gel depends on the size of the RNA, which is proportional to its charge. Specific binding of a protein to RNA alters its mobility, and the complex migrates slower compared to the free RNA. This is mainly due to an increase in the molecular mass, but also to alterations in the charge and possibly conformation. Utilizing a labeled RNA as probe allows easy monitoring of the “gel retardation” or “bandshift”. Usage of 32P-labeled RNA probes is very common and offers high sensitivity. The migration of RNA/protein complexes and free RNA are detected by autoradiography. Drawbacks are the short half-life of 32P (14.29 days), the gradual deterioration of the quality of the probe due to radiolysis, the requirement of a radioactivity license and infrastructure for radioactivity work, and potential biosafety concerns. Therefore, alternative non-isotopic methods for labeling the RNA probe have been developed, for instance with fluorophores or biotin, which enable detection by fluorescent or chemiluminescent imaging4,5. Limitations of these methods are the higher cost and often reduced sensitivity compared to isotopic labeling, and the potential of non-isotopic labels to interfere with the RNA/protein interaction. Non-denaturing polyacrylamide gels are suitable for most EMSA applications and are commonly used. On occasion, agarose gels may pose an alternative for the analysis of large complexes.
The major advantage of EMSA is that it combines simplicity, sensitivity, and robustness4. The assay can be completed within a few hours and does not require sophisticated instrumentation. RNA/protein interactions can be detected by EMSA at concentrations as low as 0.1 nM or less, and within a broad range of binding conditions (pH 4.0 – 9.5, monovalent salt concentration 1 – 300 mM, and temperature 0 – 60 °C).
RNA/protein complex formation can also be studied by the filter-binding assay. This is a simple, fast, and inexpensive procedure based on the retention of RNA/protein complexes in a nitrocellulose filter, while a free RNA probe passes through6. Compared to EMSA, it is limited in cases where the RNA probe contains multiple binding sites, or the crude extract contains more than one RNA-binding proteins that bind to the probe at the same site. While multiple RNA/protein interactions will escape detection by the filter-binding assay, they can be readily visualized by EMSA. In some cases, visualization is even possible when two RNA/protein complexes co-migrate (for instance, human IRP1/IRE and IRP2/IRE complexes), by adding an antibody against one of the RNA-binding proteins to the EMSA reaction, yielding further retardation on the gel (“supershift”)7.
The EMSA has been widely used to study IRP1 and IRP2, which are post-transcriptional regulators of iron metabolism8-10. They operate by binding to IREs, phylogenetically conserved hairpin structures within the UTRs of several mRNAs11. IREs were first discovered in the mRNAs encoding ferritin12 and transferrin receptor 1 (TfR1)13, proteins of iron storage and uptake, respectively. Later on, IREs were found in erythroid-specific aminolevulinate synthase (ALAS2)14, mitochondrial aconitase15, ferroportin16, divalent metal transporter 1 (DMT1)17, hypoxia inducible factor 2α (HIF2α)18, and other mRNAs19-21. The prototype H- and L-ferritin mRNAs contain one IRE in their 5’ UTR, while TfR1 mRNA contains multiple IREs in its 3’ UTR. IRE/IRP interactions specifically inhibit ferritin mRNA translation by sterically blocking its association of the 43S ribosomal subunit; moreover, they stabilize TfR1 mRNA against endonucleolytic cleavage. IRP1 and IRP2 share extensive sequence similarity and exhibit high IRE-binding activity in iron-starved cells. In iron-replete cells, IRP1 assembles a cubane Fe-S cluster that converts it to cytosolic aconitase at the expense of its IRE-binding activity, while IRP2 undergoes proteasomal degradation. Thus, the IRE/IRP interactions depend upon the cellular iron status, but are also regulated by other signals, such as H2O2, nitric oxide (NO) or hypoxia. Here, we describe the protocol for assessing IRE-binding activity from crude cell and tissue extracts by EMSA. We used a 32P-labeled H-ferritin IRE probe that was generated by in vitro transcription from a plasmid DNA template (I-12.CAT), where the IRE sequence was originally introduced in sense orientation downstream of the T7 RNA polymerase site by cloning of annealed synthetic oligonucleotides 22.
Protocol
Experimental procedures with mice were approved by the Animal Care Committee of McGill University (protocol 4966).
1. Preparation of Protein Extracts from Cultured Cells
- Wash cultured cells twice with 10 ml of ice-cold phosphate buffered saline (PBS).
- Scrape adherent cells with either a rubber policeman or a plastic cell scraper in 1 ml of ice-cold PBS, transfer suspension into a 1.5 ml microcentrifuge tube.
- Spin in a microcentrifuge for 5 min at 700 x g, at 4 °C. Aspirate PBS.
- Add 100 μl of ice-cold cytoplasmic lysis buffer (Table 1) per 107 cells, and pipette up and down.
- Incubate on ice for 20 min.
- Spin for 10 min at full speed in a microcentrifuge at 4 °C.
- Discard pellet. Transfer supernatant into new 1.5 ml microcentrifuge tube and keep on ice.
- Determine protein concentration (usually 1 – 10 μg/μl) using the Bradford assay23.
- Aliquot and store cell extracts at -80 °C until use.
2. Preparation of Protein Extracts from Mouse Liver and Spleen
- Euthanize a mouse with CO2 inhalation.
- Lay the euthanized animal on a clean pad over a dissecting board. Open the abdomen with scissors.
- Dissect the liver and the spleen by using scissors and forceps, and rinse each tissue in approximately 50 ml ice-cold PBS.
- Immediately cut tissues into small pieces with a scalpel (for example: approximately 1 – 2 mm3).
- Without delay, put pieces of tissues in a fresh cryotube and then snap freeze them in liquid nitrogen. Store snap-frozen tissue aliquots at -80 °C until use.
- Homogenize one piece of frozen tissue (approximately 1 – 2 mm3) in 0.25 – 0.5 ml of ice-cold cytoplasmic lysis buffer (Table 1) with a tissue homogenizer for 10 sec.
- Transfer homogenate to 1.5 ml microcentrifuge tube and chill on ice for 20 min.
- Spin for 10 min at full speed in a microcentrifuge at 4 °C.
- Discard pellet and transfer supernatant into new 1.5 ml microcentrifuge tube. Keep on ice.
- Determine protein concentration (usually 1 – 10 μg/μl) using the Bradford assay23.
- Aliquot and store cell extracts at -80 °C until use.
3. Preparation of Radiolabeled IRE-probe
- Linearize the IRE-containing plasmid I-12.CAT22 by incubating at 37 °C for 1 hr with the restriction endonuclease XbaI (1 U per μg of plasmid), which cleaves downstream of the IRE sequence. The linearized plasmid will be used as template for in vitro transcription.
- Set up an in vitro transcription reaction in a total volume of 20 μl. Use the stock solutions shown in Table 2 and add: 1 μl linearized plasmid template, 4 μl transcription buffer; 1 μl mix of ATP/CTP/GTP mix, 10 μl [α-32P]-UTP, 2 μl dithiothreitol, 1 μl RNase inhibitor and 1 μl T7 RNA polymerase. Mix by pipetting up and down.
- Incubate at 40 °C for 1 hr24.
4. Purification of Radiolabeled IRE-probe
- Terminate in vitro transcription reaction by adding 1 μl of 0.5 M EDTA, pH 8. Mix by pipetting up and down.
- Add 10 μl of 10 mg/ml tRNA, as carrier for better precipitation. Mix by pipetting up and down.
- Add 82.5 μl of 3 M ammonium acetate. Mix by vortexing.
- Add 273 μl of ethanol. Mix by vortexing.
- Let stand at RT for 5 min.
- Spin for 10 min at full speed in a microcentrifuge at RT. Discard supernatant.
- Wash pellet with 100 μl of 70% ethanol.
- Spin for 10 min at full speed in a microcentrifuge at RT. Discard supernatant.
- Air dry pellet for 10 min.
- Resuspend pellet in 100 μl of double distilled, previously autoclaved H2O.
- Quantify radioactivity in a liquid scintillation counter, aliquot radiolabeled IRE probe and store at -80 °C until use. Frozen aliquots can be used for up to 3 weeks.
5. Preparation of a native polyacrylamide gel for EMSA
- Assemble the gel (16 x 16 cm) by using 1.5 mm spacers and comb.
- To prepare a 6% native polyacrylamide gel, use the stock solutions shown in Table 3. Mix 7.5 ml of 40% acrylamide:bisacrylamide, 5 ml of 5x TBE and 37.5 ml double distilled H2O.
- Add 0.5 ml of 10% freshly prepared ammonium persulfate (APS) and 25 μl tetramethylethylenediamine (TEMED).
- Immediately pour the acrylamide solution to the gel and let it polymerize. Wait for approximately 30 min.
- Assemble the electrophoresis apparatus with the gel, fill the tanks with 0.5x TBE and connect with the power supply.
6. Electrophoretic Mobility Shift Assay
- Dilute 25 μg of protein extract from cells or tissues with cytoplasmic lysis buffer (Table 1) in a total volume of in 10 μl (lower protein concentrations may also be used). Keep on ice. If necessary, add 1 μl of 1:4 diluted 2-mercaptoethanol (2-ME) to activate dormant IRP1 (final concentration: 2%)25.
- Dilute the radiolabeled IRE probe in double distilled H2O to 200,000 cpm/μl, heat denature at 95 °C for 1 min, and cool down at RT for at least 5 min.
- Set up an EMSA reaction by adding 1 μl of radiolabeled IRE probe to the protein extract.
- Incubate for 20 min at RT.
- Add 1 μl of 50 mg/ml heparin to the reaction (to inhibit non-specific protein interactions with the probe9) and continue the incubation for another 10 min. Aliquot the stock solution of heparin (50 mg/ml) and store at -80 °C.
- When using long radiolabeled probes (>60 nucleotides), add 1 μl RNase T1 (1 U/μl) and incubate for 10 min at RT to reduce non-specific protein binding to the probe, and to allow better separation of the RNA/protein complex during electrophoresis.
- Add 3 μl of loading buffer (80% glycerol + bromophenol blue), mix and load on the 6% non-denaturing polyacrylamide gel.
- Run the gel for 60 min at 130 V (5 V/cm).
- Transfer the gel onto a large filter paper and dry.
- Expose to a film and develop autoradiography. Exposure time may range from 1 hr (or less) up to O/N.
Representative Results
A radiolabeled IRE probe was prepared, as described in sections 3 and 4 of the protocol. The sequence of the probe was 5'-GGGCGAAUUC GAGCUCGGUA CCCGGGGAUC CUGCUUCAAC AGUGCUUGGA CGGAUCCU-3'; the bolded nucleotides represent an unpaired C residue and the loop, which are critical IRE features. The specific radioactivity of the probe was 4.5 x 109 cpm/μg of RNA.
To assess the effects of iron perturbations on IRE-binding activity, murine RAW264.7 macrophages were left untreated, or treated with hemin (iron source) or desferrioxamine (iron chelator). Lysates were prepared and analyzed by EMSA with the IRE probe (8,000 cpm/μg protein). Representative data are shown in Figure 1, with shorter and longer exposures of the autoradiograms on the left and right panels, respectively. Murine IRE/IRP1 and IRE/IRP2 complexes exhibit different mobility and two distinct bands are visible in lane 1, corresponding to untreated cells. Iron chelation profoundly induced the IRE-binding activities of both IRP1 and IRP2 (lane 2), while iron supplementation diminished them (lane 3). Pretreatment with 2% 2-ME fully activated IRP1 (bottom panel), even in extracts of iron-treated cells. This demonstrates that iron perturbations do not impinge on the stability of IRP1 and also serves as a loading control. By contrast the 2-ME pretreatment inhibited the IRE-binding activity of IRP2. The fast migrating non-specific bands are not affected by iron.
Next, we analyzed IRE-binding activity in livers of wild type, Irp1-/- and Irp2-/- mice, previously fed for one week with a standard or an iron-enriched diet (Figure 2). The Irp1-/- and Irp2-/- mice were kindly provided by Dr. M. W. Hentze (EMBL, Heidelberg). In wild type liver extracts, IRP1 accounted for the major fraction of IRE-binding activity and IRE/IRP2 complexes were hardly visible (lanes 1, 2), in agreement with previous observations26. IRP2 exhibited potent IRE-binding activity in liver extracts of Irp1-/- mice (lanes 3, 4). Under these conditions, no IRE/IRP1 complexes were observed. Likewise, no IRE/IRP2 complexes were formed in liver extracts of Irp2-/- mice (lanes 5, 6). Feeding mice with a high-iron diet decreased hepatic IRE-binding activity of both IRP1 and IRP2; this effect was more dramatic on IRP2 (lanes 7-12). Note that after treatment of wild type or Irp2-/- liver extracts with 2-ME, the IRE-binding activity of IRP1 was enhanced to a point where it shifted almost all IRE probe (this is clearly observed in the shorter exposure of the autoradiograms on the left panel).
Finally, we evaluated the IRE-binding activity in livers and spleens of wild type and Hjv-/- mice (Figure 3). The Hjv-/- mice were kindly provided by Dr. N. C. Andrews (Duke University, NC). These animals represent a model of hereditary hemochromatosis27, a disease of systemic iron overload, where excessive iron accumulates in parenchymal cells, while reticuloendothelial macrophages remain iron deficient28. As expected, livers of Hjv-/- mice (with iron-loaded hepatocytes) exhibited decreased IRE-binding activity compared to wild type (lanes 1-4). Conversely, IRE-binding activity was higher in spleens of Hjv-/- mice (containing iron-deficient macrophages; lanes 5-8). Again here, the 2-ME treatment promoted almost complete shift of the IRE probe in liver extracts (see shorter exposure of the autoradiograms on the left panel).
Table 1. Cytoplasmic lysis buffer.
| 1% Triton X100 |
| 25 mM Tris-HCl, pH 7.4 |
| 40 mM KCl |
- The solution should be autoclaved and stored at RT.
- Protease inhibitors, such as 10 μg/ml leupeptin and 0.1 mM phenylmethanesulfonyl fluoride (PMSF) may be added prior to use.
- Addition of fresh 1 mM dithiotheritol is recomended for detection of IRP2 activity.
Table 2. Stock solutions for in vitro transcription reaction.
| 1 μg/μl linearized plasmid template |
| 5x transcription buffer (supplied with T7 RNA polymerase) |
| 20 mM mix of ATP, CTP and GTP |
| 3,000 Ci/mmol [α-32P]-UTP |
| 100 mM dithiothreitol |
| 10 U/μl RNase inhibitor |
| 20 U/μl T7 RNA polymerase |
Table 3. Stock solutions for non-denaturing polyacrylamide gel electrophoresis.
| 40% acrylamide:bisacrylamide (37.5:1) |
| 5x TBE (Tris/borate/EDTA) |
For making 1 L of 5x TBE, use 54 g of Tris base, 27.5 g of boric acid, and 20 ml of 0.5M EDTA (pH 8.0).
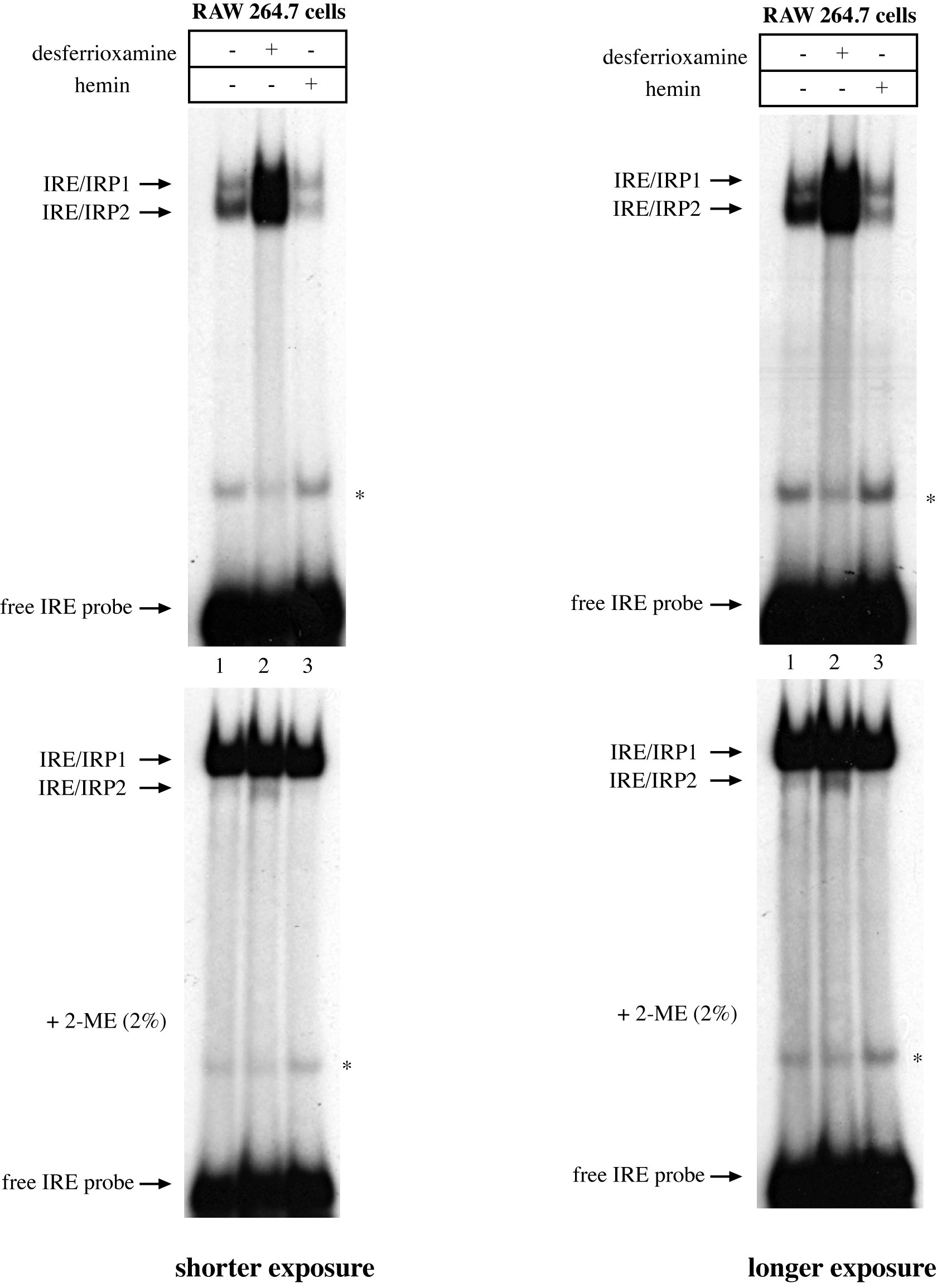
Figure 1. Iron-dependent regulation of IRE-binding activities in RAW264.7 macrophages. 107 cells were either left untreated or treated O/N with 100 μM hemin or desferrioxamine. Cytoplasmic lysates were prepared and analyzed by EMSA with a 32P-labeled IRE probe in the absence (top) or presence of 2% 2-ME (bottom). The positions of IRE/IRP1 and IRE/IRP2 complexes, and of free IRE probe are indicated by arrows. The star indicates a non-specific band. Shorter and longer exposures of the autoradiograms are shown in the left and right panels, respectively. Please click here to view a larger version of this figure.
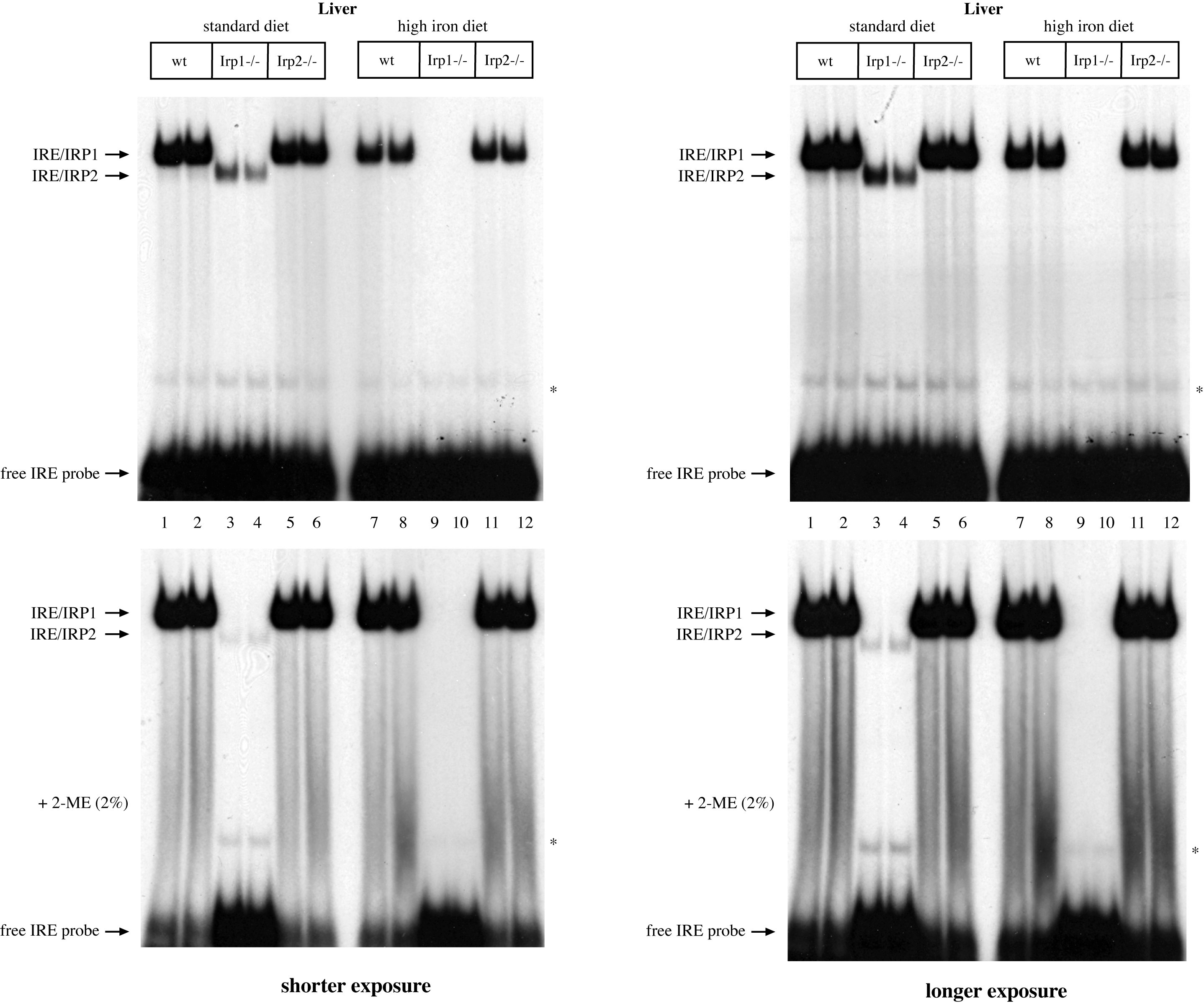
Figure 2. Analysis of IRE-binding activities in livers of wild type (wt), Irp1-/- and Irp2-/- mice. 5 week old mice (n = 2 for each genotype), all in C57BL/6 background29, were placed on a standard or a high-iron diet (containing 2% carbonyl iron). After one week the animals were euthanized. Hepatic protein extracts were prepared and analyzed by EMSA with a 32P-labeled IRE probe in the absence (top) or presence of 2% 2-ME (bottom). The positions of IRE/IRP1 and IRE/IRP2 complexes, and of free IRE probe are indicated by arrows. The star indicates a non-specific band. Shorter and longer exposures of the autoradiograms are shown in the left and right panels, respectively. Please click here to view a larger version of this figure.

Figure 3. Analysis of IRE-binding activities in livers and spleens of wild type (wt) and Hjv-/- mice. 10 week old mice (n = 2 for each genotype), all in C57BL/6 background30, were euthanized. Hepatic and splenic protein extracts were prepared and analyzed by EMSA with a 32P-labeled IRE probe in the absence (top) or presence of 2% 2-ME (bottom). The positions of IRE/IRP1 and IRE/IRP2 complexes, and of free IRE probe are indicated by arrows. The star indicates a non-specific band. Shorter and longer exposures of the autoradiograms are shown in the left and right panels, respectively. Please click here to view a larger version of this figure.
Discussion
Herein, we describe a protocol that has been developed to study the IRE-binding activities of IRP1 and IRP2, and we show representative data. By using different probes, this protocol can also be adjusted for the study of other RNA-binding proteins. A critical step is the size of the probe. Usage of long probes, which is common when the exact binding site is unknown, can result in RNA/protein complexes that do not migrate differently than the free RNA. In this case, it is advisable to remove unbound RNA by treatment with RNAse T1 (step 6.6). The quality of the probe is also very important. Best results are obtained with freshly prepared radiolabeled RNA. Gel purification10 is not necessary, but may improve the appearance of the EMSA if the in vitro transcription reaction for probe preparation yields premature termination products (and if no RNAse T1 digestion step is included). Running the native gel electrophoresis too fast may result in overheating, which could lead to dissociation of RNA/protein complexes and result in fuzzy bands.
Quantitative results can be obtained by analyzing the intensity of the retarded bands with phosphorimaging. Quantifications are only valid when the RNA probe is present in excess and is not limiting for RNA binding. The affinity of the RNA/protein interaction, represented by the dissociation constant Kd that is linked to the free energy ΔGo, can be assessed under equilibrium conditions by titrating the RNA-binding activity with decreasing amounts of radiolabeled probe3. The IRE/IRP1 interaction has been extensively characterized physicochemically9,31. Titration EMSA experiments with varying amounts of an RNA-binding protein can be performed to calculate stoichiometry3. Specificity for binding can be easily validated by competition experiments with excess of unlabeled “cold” competitor RNAs3. Only specific competitors should reduce the intensity of the retarded radioactive band. Specificity can also be monitored by including an antibody against the RNA-binding protein in the EMSA reaction, which will yield a “supershift”7. This can also be exploited to determine the identity of the RNA-binding protein. No “supershift” should be observed with non-specific antibodies or pre-immune serum.
In conclusion, the EMSA is a powerful method for the analysis of RNA/protein interactions. It is relatively easy to perform and requires minimal infrastructure. It is not as simple as the filter-binding assay, but will yield more informative and accurate data when multiple RNA-binding proteins interact specifically with the probe (such as IRP1 and IRP2). It should be noted that unlike murine, human IRE/IRP1 and IRE/IRP2 complexes co-migrate in EMSA. However, they can be separated by “supershifts” with specific antibodies7. The EMSA is not valuable for the detailed mapping of RNA/protein interactions; other techniques, such as the RNAse protection assay, toeprinting or cross-linking32 are more appropriate. Nevertheless the EMSA is an excellent choice for discovering new RNA/protein interactions, as has been the case with IRPs8, but also for corroborating data obtained with computational or high throughput experimental approaches. Moreover, the EMSA is superior for analyzing specificity and physicochemical properties of RNA/protein interactions.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was supported by a grant from the Canadian Institutes for Health Research (MOP-86514).
Materials
| Name of the Materials | Company | Catalog Number |
| leupeptin | SIGMA | L2884 |
| PMSF | SIGMA | 78830 |
| BioRad Protein Assay | BIORAD | 500-0006 |
| T7 RNA polymerase | Thermoscientific | EPO111 |
| Rnase Inhibitor | Invitrogen | 15518-012 |
| UTP [alpha-32P] | Perkin-Elmer | NEG507H |
| Scintillation liquid | Beckman Coulter | 141349 |
| heparin | SIGMA | H0777 |
| Rnase T1 | Thermoscientific | EN0541 |
| Name of the Equipments | Company | Catalog Number |
| Tissue Ruptor | Qiagen | 9001271 |
| Scintillation counter | Beckman Coulter | LS6500 |
| Protean II xi Cell | BIORAD | 165-1834 |
| 20 wells combs 1.5mm thick | BIORAD | 165-1868 |
| 1.5 mm spacers | BIORAD | 165-1849 |
| PowerPac | BIORAD | 164-5070 |
References
- Fried, M., Crothers, D. M. Equilibria and kinetics of lac repressor-operator interactions by polyacrylamide gel electrophoresis. Nucleic Acids Res. 9, 6505-6525 (1981).
- Garner, M. M., Revzin, A. A gel electrophoresis method for quantifying the binding of proteins to specific DNA regions: application to components of the Escherichia coli lactose operon regulatory system. Nucleic Acids Res. 9, 3047-3060 (1981).
- Ryder, S. P., Recht, M. I., Williamson, J. R. Quantitative analysis of protein-RNA interactions by gel mobility shift. Methods Mol Biol. 488, 99-115 (2008).
- Hellman, L. M., Fried, M. G. Electrophoretic mobility shift assay (EMSA) for detecting protein-nucleic acid interactions. Nat Protoc. 2, 1849-1861 (2007).
- Luscieti, S., et al. Novel mutations in the ferritin-L iron-responsive element that only mildly impair IRP binding cause hereditary hyperferritinaemia cataract syndrome. Orphanet J Rare Dis. 8, 30 (2013).
- Rio, D. C. . Filter-binding assay for analysis of RNA-protein interactions. 2012, 1078-1081 (2012).
- Wang, J., et al. Iron-mediated degradation of IRP2: an unexpected pathway involving a 2-oxoglutarate-dependent oxygenase activity. Mol. Cell. Biol. 24, 954-965 (2004).
- Leibold, E. A., Munro, H. N. Cytoplasmic protein binds in vitro to a highly conserved sequence in the 5′ untranslated regions of ferritin heavy- and light-subunit mRNAs. Proc. Natl. Acad. Sci. USA. 85, 2171-2175 (1988).
- Haile, D. J., Hentze, M. W., Rouault, T. A., Harford, J. B., Klausner, R. D. Regulation of interaction of the iron-responsive element binding protein with iron-responsive RNA elements. Mol. Cell. Biol. 9, 5055-5061 (1989).
- Mueller, S., Pantopoulos, K. Activation of iron regulatory protein-1 (IRP1) by oxidative stress. Methods Enzymol. 348, 324-337 (2002).
- Wang, J., Pantopoulos, K. Regulation of cellular iron metabolism. Biochem J. 434, 365-381 (2011).
- Hentze, M. W., et al. Identification of the iron-responsive element for the translational regulation of human ferritin mRNA. Science. 238, 1570-1573 (1987).
- Casey, J. L., et al. Iron-responsive elements: regulatory RNA sequences that control mRNA levels and translation. Science. 240, 924-928 (1988).
- Dandekar, T., et al. Identification of a novel iron-responsive element in murine and human erythroid d-aminolevulinic acid synthase mRNA. EMBO J. 10, 1903-1909 (1991).
- Gray, N. K., Pantopoulos, K., Dandekar, T., Ackrell, B. A. C., Hentze, M. W. Translational regulation of mammalian and drosophila citric acid cycle enzymes via iron-responsive elements. Proc. Natl. Acad. Sci. USA. 93, 4925-4930 (1996).
- McKie, A. T., et al. A novel duodenal iron-regulated transporter IREG1, implicated in the basolateral transfer of iron to the circulation. Mol. Cell. 5, 299-309 (2000).
- Gunshin, H., et al. Cloning and characterization of a mammalian protein-coupled metal-ion transporter. Nature. 388, 482-488 (1997).
- Sanchez, M., Galy, B., Muckenthaler, M. U., Hentze, M. W. Iron-regulatory proteins limit hypoxia-inducible factor-2alpha expression in iron deficiency. Nat. Struct. Mol. Biol. 14, 420-426 (2007).
- Sanchez, M., et al. Iron regulation and the cell cycle: Identification of an iron-responsive element in the 3′-untranslated region of human cell division cycle 14A mRNA by a refined microarray-based screening strategy. J. Biol. Chem. 281, 22865-22874 (2006).
- Santos, C. O., et al. An iron responsive element-like stem-loop regulates alpha-hemoglobin-stabilizing protein mRNA. J Biol Chem. 283, 26956-26964 (2008).
- Liu, Z., et al. Siderophore-mediated iron trafficking in humans is regulated by iron. J Mol Med (Berl. 90, 1209-1221 (2012).
- Gray, N. K., et al. Recombinant iron regulatory factor functions as an iron-responsive element-binding protein, a translational repressor and an aconitase. A functional assay for translational repression and direct demonstration of the iron switch. Eur. J. Biochem. 218, 657-667 (1993).
- Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248-254 (1976).
- Testa, U., et al. Differential regulation of iron regulatory element-binding protein(s) in cell extracts of activated lymphocytes versus monocytes-macrophages. J. Biol. Chem. 266, 13925-13930 (1991).
- Hentze, M. W., Rouault, T. A., Harford, J. B., Klausner, R. D. Oxidation-reduction and the molecular mechanism of a regulated RNA-protein interaction. Science. 244, 357-359 (1989).
- Meyron-Holtz, E. G., et al. Genetic ablations of iron regulatory proteins 1 and 2 reveal why iron regulatory protein 2 dominates iron homeostasis. EMBO J. 23, 386-395 (2004).
- Huang, F. W., Pinkus, J. L., Pinkus, G. S., Fleming, M. D., Andrews, N. C. A mouse model of juvenile hemochromatosis. J. Clin. Invest. 115, 2187-2191 (2005).
- Sebastiani, G., Pantopoulos, K. Disorders associated with systemic or local iron overload: from pathophysiology to clinical practice. Metallomics. 3, 971-986 (2011).
- Galy, B., Ferring, D., Hentze, M. W. Generation of conditional alleles of the murine iron regulatory protein (IRP)-1 and -2 genes. Genesis. 43, 181-188 (2005).
- Gkouvatsos, K., et al. Iron-dependent regulation of hepcidin in Hjv-/- mice: Evidence that hemojuvelin is dispensable for sensing body iron levels. PLoS ONE. 9, 85530 (2014).
- Goforth, J. B., Anderson, S. A., Nizzi, C. P., Eisenstein, R. S. Multiple determinants within iron-responsive elements dictate iron regulatory protein binding and regulatory hierarchy. RNA. 16, 154-169 (2010).
- Gopinath, S. C. Mapping of RNA-protein interactions. Anal Chim Acta. 636, 117-128 (2009).

