Split-and-pool Synthesis and Characterization of Peptide Tertiary Amide Library
Summary
Peptide tertiary amides (PTAs) are a superfamily of peptidomimetics that include but are not limited to peptides, peptoids and N-methylated peptides. Here we describe a synthetic method which combines both split-and-pool and sub-monomer strategies to synthesize a one-bead one-compound library of PTAs.
Abstract
Peptidomimetics are great sources of protein ligands. The oligomeric nature of these compounds enables us to access large synthetic libraries on solid phase by using combinatorial chemistry. One of the most well studied classes of peptidomimetics is peptoids. Peptoids are easy to synthesize and have been shown to be proteolysis-resistant and cell-permeable. Over the past decade, many useful protein ligands have been identified through screening of peptoid libraries. However, most of the ligands identified from peptoid libraries do not display high affinity, with rare exceptions. This may be due, in part, to the lack of chiral centers and conformational constraints in peptoid molecules. Recently, we described a new synthetic route to access peptide tertiary amides (PTAs). PTAs are a superfamily of peptidomimetics that include but are not limited to peptides, peptoids and N-methylated peptides. With side chains on both α-carbon and main chain nitrogen atoms, the conformation of these molecules are greatly constrained by sterical hindrance and allylic 1,3 strain. (Figure 1) Our study suggests that these PTA molecules are highly structured in solution and can be used to identify protein ligands. We believe that these molecules can be a future source of high-affinity protein ligands. Here we describe the synthetic method combining the power of both split-and-pool and sub-monomer strategies to synthesize a sample one-bead one-compound (OBOC) library of PTAs.
Introduction
Peptidomimetics are compounds that mimic the structure of natural peptides. They are designed to retain the bioactivity while overcoming some of the problems associated with natural peptides, including cell permeability and stability against proteolysis1-3. Due to the oligomeric nature of these compounds, large synthetic libraries can be readily accessed through monomeric or sub-monomeric synthetic routes4-7. One of the most studied classes of peptidomimetics is peptoids. Peptoids are oligomers of N-alkylated glycines that can be synthesized easily using a sub-monomer strategy8,9. Many useful protein ligands have been successfully identified from screening large synthetic peptoid libraries against protein targets1,10-14. Nonetheless, “hits” identified from peptoid libraries rarely archive very high affinity towards protein targets1,10-14,22. One major difference between peptoids and natural peptides is that most of the peptoids generally lack the ability to form secondary structure due to the lack of chiral centers and conformational constraints. In order to solve this problem, multiple strategies were developed over the past decade, largely focusing on the modification of side chains contained on the main chain nitrogen atoms15-22. Recently, we have developed a new synthetic route to introduce natural amino acid side chains onto a peptoid backbone to create peptide tertiary amides23.
Peptide tertiary amides (PTAs) are a super family of peptidomimetics that include but are not limited to peptides (R2 = H), peptoids (R1 = H) and N-methylated peptides (R1 ≠ H, R2 = Me). (See Figure 1) Our synthetic route employs naturally occurring amino acids as the source of chirality and side chains on the α-carbon, and commercially available primary amines to provide N-substitutions. Therefore, a larger chemical space than that of simple peptides, peptoids or N-methylated peptides can be explored. Circular dichroism spectra have shown that PTA molecules are highly structured in solution. Characterization of one of the PTA-protein complexes clearly shows that the conformational constraints of PTA are required for binding. Recently, we have also discovered that some of the PTA molecules possess improved cell permeability than their peptoid and peptide counterparts. We believe that these PTA libraries can be a good source of high-affinity ligands for protein targets. In this paper, we will discuss the synthesis of a sample one-bead one-compound (OBOC) PTA library in details along with some improved conditions for the coupling and cleavage of these compounds.
Protocol
1. Basics of Split-and-pool Synthesis
In order to efficiently generate a large number of compounds on solid phase, split-and-pool synthesis is often employed as a general strategy. As shown in Figure 4, tentagel beads are first split into three portions. Each portion is reacted with a different reagent, generating the first residue on beads. After the first reaction, all three portions are pooled together, mixed, and then split again into three portions. Each portion will again react with a different reagent, generating the second residue on beads. After two split-and-pool steps, nine compounds are generated.
In sub-monomer synthesis, beads are first divided into several portions to react with different bromo acids in the presence of coupling reagent. After washing with solvent, all beads will be pooled together and mixed, then again divided into several portions to react with different primary amines. After amination, all beads are pooled together and washed thoroughly, completing a full monomer on each bead. This process can be repeated till desired diversity is reached.
2. Preparation of Acid Bromide from Natural Amino Acids
In sub-monomer synthesis, the synthesis of each monomer is divided into two separate steps: 1. coupling of acid bromide and 2. amination with primary amines (Figure 2). In order to synthesize a peptide tertiary amide, chiral acid bromides with side chains on the alpha carbon will be prepared from natural amino acids. Here we describe the method of transforming a natural amino acid into the corresponding acid bromide with high stereo fidelity. We use alanine as an example; other amino acids including serine, threonine, aspartic acid, glutamic acid, asparagine, glutamine, glycine, valine, isoleucine, phenylalanine can also be transformed into bromo acids under similar conditions. Note that some of the amino acids with functional groups like phenol, guanidine and amine need to be protected before the transformation. The reaction setup is shown in Figure 3.
Safety Precaution: For the following reactions involving HBr, NaNO2 and other corrosive/toxic chemicals, proper safety equipment like safety goggles, lab coat, and chemical resistant gloves are needed. All reactions should be performed in a fume hood by experienced chemist.
- Add 370 ml water into 630 ml 48% HBr solution to prepare a 1 L, 30% HBr solution. Add 500 ml ethylene glycol into a 1 L bath container; add dry ice to maintain the temperature at -10 °C. Caution: 48% HBr solution is strongly acidic and corrosive, handle with care. Read the MSDS before use.
- Add D-alanine (8.9 g, 0.1 mol) and KBr (11.9 g, 0.1 mol) to a 250 ml three-neck round bottom flask with a magnetic stir bar. Add 100 ml, and the 30% HBr prepared in the previous step. Put the flask in the ethylene glycol bath prepared in step 2.1 and keep the temperature at -10 °C. Bubble argon through a long needle from the bottom of the flask for 10 min as shown in Figure 3. Stir the solution with the magnetic stir bar at 300 rpm.
- Dissolve NaNO2 (8.28 g, 0.12 mol) in a 100 ml beaker with 20 ml water. Add the solution into the pressure equalizing dropping funnel and seal the dropping funnel with a septum. Slowly turn on the valve of the dropping funnel and let the NaNO2 solution drop into the flask. Control the valve to adjust the dripping rate to approximately 2 drops per sec. Keep the stirring at 300 rpm and keep the argon bubbling from the bottom of the flask. The flask should be kept in ethylene glycol bath at -10 °C until all NaNO2 is added. Caution: This step generates heat and gas during the addition of NaNO2 solution. Dripping rate should be carefully controlled and the whole system should be open through the argon outlet.
- Keep stirring for 3 more hr and let the temperature warm up from -10 °C to room temperature. The resulting solution should be clear to light yellow; if the color is too dark, apply vacuum to remove the excess nitrogen oxides and possible Br2 generated during the reaction.
- Extract the product from the solution with 3x 35 ml diethyl ether using an extracting funnel. Combine the organic phase and wash it with saturated brine. The organic phase can also be washed with a small amount of NaHCO3 prior to washing with brine to remove the color if it is dark. Dry the organic phase over Na2SO4 for 6 hr.
- Filter out the Na2SO4 and evaporate the solvent under vacuum, crude product should be obtained as clear to pale yellow oil. Crude product can be further purified by distillation at 115 °C, 3 mm Hg, or by silica column with 3:1 hexane:ethyl acetate.
- Pure product is obtained as clear oil, 6.6 g (yield 74%), density = 1.69 g/ml, [α]D20 = +24° (methanol), 1H NMR (400 MHz, CDCl3) δ 4.41 (q, J = 7.0 Hz, 1H), 1.86 (d, J = 7.0 Hz, 3H). In the case of (S)-2-bromopropanoic acid-d4 (prepared from d4-L-alanine), pure product is obtained as clear oil, yield 78%, density = 1.72 g/ml, [α]D20 = -19° (methanol). 1H NMR, no significant H signal is observed. ESI-MS– [M-1]- = 155.1 (expected 154.97). For (S)-2-bromo-4-methylpentanoic acid (prepared from L-leucine using the same procedure) pure product is obtained as clear oil, yield 89%, [α]D20 = +37° (methanol), 1H NMR (400 MHz, CDCl3) δ 4.30 (t, J = 7.7 Hz, 1H), 1.94 (dd, J = 10.8, 3.9 Hz, 2H), 1.81 (tt, J = 13.2, 6.5 Hz, 1H), 0.96 (dd, J = 18.2, 6.6 Hz, 7H). In the case of (S)-2-bromo-3-phenylpropanoic acid (prepared from L-phenylalanine using the same procedure) pure product is obtained as pale yellow oil, yield 72%, [α]D20 = +17° (methanol), 1H NMR (400 MHz, CDCl3) δ 7.38 – 7.19 (m, 5H), 4.42 (dd, J = 8.1, 7.3 Hz, 1H), 3.47 (dd, J = 14.2, 8.2 Hz, 1H), 3.25 (dd, J = 14.2, 7.2 Hz, 1H).
3. Isotopic Labeling of Alanine Using Transaminase
In combinatorial library synthesis, especially in the split-and-pool synthesis of one-bead one-compound (OBOC) libraries, the amount of compound that can be obtained from each bead is relatively small. (Typically 1 pmol to 10 nmol). Additionally, mass spectrometry is widely used for the identification and characterization of the final compound due to its high sensitivity. In order to use mass spectrometry to determine the absolute stereochemistry at the chiral centers of the final PTA products, bromo acid enantiomers should be isotopically labeled before use. Here we describe the method of using transaminase and D2O to label L-alanine.
- Dissolve L-alanine (300 mg, 3.36 mmol) with 10 ml of D2O in a 50 ml polyethylene tube. Add α-ketoglutarate (10 mg, 0.068 mmol) as co-substrate. Warm up the tube to 37 °C and adjust the pD to 8.5-8.7 using a 1 M NaOD soution. Note: pD is determined by pH test strips. Traditional electro pH meter equipped with glass electrode selective for H+ may give incorrect read out for D+.
- Add alanine transaminase (0.1 mg, EC 2.6.1.2 from pig heart, Roche Diagnostics, Indianapolis, IN) to the pD 8.5 – 8.7, 37 °C solution prepared from previous step. Put the tube in a 37 °C incubator and incubate it overnight with mild shaking, 10 to 30 rpm is preferred.
- After overnight incubation, take 0.5 ml of the D2O solution and check the reaction progress by 1H-NMR. All proton signals of alanine, δ 3.76 (q, J = 7.2 Hz, 1H), 1.46 (d, J = 7.3 Hz, 3H), 1H NMR 400 MHz, should be greatly suppressed due to deuteration. More than 98% of the proton should be exchanged to deuterium as described previously23. Note: D2O can be partially recovered by distillation if the reaction is performed on large scale (>200 ml D2O). Normally, 60% to 80% of D2O can be distilled from the solution.
- Freeze the above solution with liquid nitrogen and lyophilize it using a lyophilizer to obtain white deuterated L-alanine powder.
4. Synthesis of Peptoid Linker Region
The linker region is not required for PTA library synthesis. However, in order to avoid the high background in the lower molecular weight range (100-600) of MALDI mass spectroscopy and to improve the ionization of the compounds, a peptoid linker with multiple polar residues is often used. This peptoid linker can be synthesized through standard peptoid synthesis procedure. Here we will synthesize a pentamer of N-methoxyethyl glycine as the linker (as shown in Figure 5).
- Swell 90 μm Tentagel beads with RAM linker (1 g, 0.27 mmol/g) in 10 ml DMF for 3 hr in a 12 ml syringe reactor with mild shaking.
- Drain the DMF from the reactor and add 10 ml 20% piperidine DMF solution to deprotect the fmoc group from the Rink amide linker. Shake the beads with 20% piperidine solution for 30 min. Wash with DMF 5x to remove all piperidine.
- Take a few beads out from the syringe and test it with chloranil test. Beads should turn dark brown (chloranil test positive for primary amine) if fmoc is successfully deprotected.
- Prepare the following solutions: 1. 20 ml, 2 M bromoacetic acid/DMF solution; 2. 20 ml, 2 M DIC/DMF solution; 3. 10 ml, 1 M methoxylethylamine/DMF solution.
- Add 5 ml of 2 M bromoacetic acid/DMF solution to the beads, shake gently. Then add 5 ml of 2 M DIC/DMF solution to the beads; seal the syringe with the plunger and put it on the shaker. Shake for 10 min.
- Wash the beads with DMF thoroughly. Add 2 ml of 1 M methoxyethylamine/DMF solution prepared from step 4.4 to the beads. Seal the syringe with the plunger and shake it on the shaker for 30 min.
- Wash the beads with DMF 5x. Check a few beads with chloranil test, if positive (beads turn blue), then continue to the next step. Otherwise, repeat step 4.6.
- Repeat steps 4.5 to 4.7, 4x to complete the pentamer.
5. Split-and-pool Synthesis of PTA Library with (R)- and (S)-2-bromopropionic Acids
Here we describe the synthesis of a small PTA library with a theoretical diversity of 9,261 compounds using the 1 g of beads from step 4.8. Note that a 90 µm tentagel bead contains approximately 2.9 million beads per gram; therefore the redundancy of the library will be 2.9 x 106 / 9,261 = 312 copies. We will use bromoacetic acid, (R)-2-bromopropanoic and isotopic labeled (S)-2-bromopropanoic acid-d4 as the acids, and 7 different amines (A1 ~ A7, see Figure 5 for details) for amination. Syringe reactors and a vacuum manifold will be used to perform the synthesis.
- Add 10 ml of 1:1 DCM:DMF to the syringe from step 4.8; use a 1,000 μl pipette with a truncated pipette tip to split all 1 g beads evenly into three 5 ml syringe reactors. Label them as B (bromoacetic acid), R ((R)-2-bromopropanoic) and S ((S)-2-bromopropanoic acid-d4). Wash all 3 syringes with DCM 3x, and wash syringes labeled with R and S with anhydrous THF 3x, wash syringe labeled with B with DMF 3x.
- Syringe R and S. BTC coupling of bromopropanoic acid.
- Prepare a fresh BTC/THF solution. Add approximately 200 mg of BTC into the vial in a fume hood, seal it with the cap. Weigh the amount of BTC in the vial. Calculate the amount of solvent needed and add anhydrous THF into the vial to make a 20 mg/ml BTC/THF solution.
- Prepare the bromo acids/BTC mixture. Add (R)-2-bromopropanoic acid (89 μl, 0.95 mmol) and (S)-2-bromopropanoic acid-d4 (89 μl, 0.95 mmol) in two small vials separately. To each vial, add 5 ml of the above 20 mg/ml BTC/THF solution. Seal the two vials and put them in -20 °C freezer for 20 min.
- Add 1,125 μl, 2:1 THF/DIPEA (750 μl THF, 375 μl DIPEA, 2.2 mmol) to syringe R and S separately. Mix the beads with the pipette tip. Let them sit for 5 min.
- Take the two cooled bromo acids/BTC mixtures from step 5.2.2, add 2,4,6-Trimethylpyridine (356 μl, 2.7 mmol) to each vials. White precipitates will form immediately. Apply the corresponding suspension directly to the basified beads (syringe R and S in step 5.2.3) as soon as possible and then put them on a shaker to shake under 120 rpm for 2 hr.
Note: The solution in the syringe reactors should be a pale yellowish suspension during the whole course of the reaction. A darker color is an indication of excessive heat released during the initial addition of the acid chloride solution. This can be solved by further cooling down or dilute the bromo acids/BTC mixture.
- Syringe B. Bromoacetic acid coupling with DIC
- Prepare a fresh 20 ml, 2 M bromoacetic acid/DMF solution. Prepare a 20 ml, 2 M DIC/DMF solution.
- Add 2 ml of 2 M bromoacetic acid/DMF solution to syringe B, shake gently. Add 2 ml of 2 M DIC/DMF solution to syringe B, shake gently.
- Put syringe B on the same shaker as syringe R and S, shake it for 2 hr. Note that the coupling reaction of bromoacetic acid /DIC is done within 30 min; prolonged reaction time is for the convenience of split-and-pool synthesis.
- After 2 hr, take syringe R, S and B from the shaker. Wash all three syringes thoroughly with DCM 5x. Then wash with DMF 5x. Note that syringe R and S cannot be washed with DMF before being washed with DCM or THF first.
- Pool all the beads from syringes R, S and B into one 12 ml syringe reactor. Wash all the beads with DMF 5x.
- Add 10 ml of 1:1 DCM:DMF to the syringe; use a 1,000 μl pipette with a truncated pipette tip to split all the beads evenly into 7 individual 2 ml syringes, label them as A1-A7.
- Amination. Prepare 10 ml, 2 M primary amine/DMF solutions for each of the 7 amines listed in Figure 5. Add 5 ml of each amine solution to the corresponding syringe A1-A7. Incubate all 7 syringes in a 60 °C incubator with shaking overnight.
- After incubation, wash all beads thoroughly with DMF. Take a few of beads from each syringe and check with chloranil test. If the beads turn green (positive) within 3 min, proceed with next step. If negative, repeat step 5.7 for the negative syringes.
- Repeat steps 5.1 to 5.8 2x to complete the trimer. Optional step: After each cycle, we recommend to check the synthesis quality by mass spectroscopy as described below. All 9,261 compounds are now synthesized on tentagel beads as OBOC library.
- Mass spectroscopic confirmation of PTAs.
PTAs are highly structured oligomers and possess many common features of N-methylated peptides. One of the common problems of solid phase synthesis of N-methylated peptides is the acid degradation during TFA cleavage. To suppress acid degradation, cleavage of molecules like cyclosporine from solid support is often carried out under low temperature. We compared various cleaving conditions for cleaving different PTA molecules from the solid support. We found that, generally, low temperature and reduced TFA concentration could effectively suppress acid degradation and provide purer compounds.- Prepare 10 ml 1:1 TFA/DCM solution in a 15 ml tube. Seal the tube and put it in a -20 °C freezer for 20 min.
- Wash the beads that need to be cleaved with DCM 5x. Shake the beads in DCM for 15 min and wash the beads again with DCM 5x.
- Drain the DCM from the syringe. Use a light microscope and a pipette with truncated tip to transfer each individual bead into a 96-well plate, one bead per well.
- Cover the 96-well plate with a cover slip. Put the plate in a -20 °C freezer for 15 min.
- Take the cooled 1:1 TFA/DCM solution from step 5.10.1 and add 20 μl to each of the wells that contains a bead. Put the cover slip back and put the 96-well plate on a shaker in the -20 °C fridge.
- Shake for 20 min. Take the 96-well plate out and peel off the cover slip. Blow-dry the TFA/DCM from each well by blowing air or argon over it. If more than 10 beads are cleaved, a speedvac can be used to dry the TFA/DCM from the whole plate. Note that at this point, not all of the compounds are cleaved off the beads, but the cleaved compounds should be more than enough to perform the mass spectroscopy analysis.
- Add 20 μl 6:4 ACN:H2O solution to dissolve the cleaved compounds from each well. Spot 0.6 μl of each compound solution together with 0.6 μl CHCA MALDI matrix on the MALDI plate.
- Use MALDI mass spectrometry to determine the molecular weight and sequence (MS/MS) of each compound.
6. Chloranil Test
- Prepare the following reagents fresh for each test. Solution A: 2% Chloranil (CAS: 118-75-2) in DMF. Solution B: 2% acetaldehyde (CAS: 75-07-0) in DMF.
- Mix 100 μl of solution A with 100 μl of solution B before the test in a 1.5 ml tube; drop the beads in and gently shake. If the beads turn blue within 5 min, it indicates the presence of secondary amine on the surface of the beads. Primary amines give a dark brown color instead of turning blue.
Representative Results
Here we show three representative MALDI spectrums from a PTA trimer with linker. As shown in Figure 6A, when cleaved under room temperature using 50% TFA/DCM solution, significant degradation is observed. In Figure 6A, peak 593 and 484 correspond to the linker and the PTA trimer respectively, show that the whole molecule was successfully synthesized on bead but degraded during cleavage. When cleaved under low temperature condition as described above, the amount of TFA-induced degradation is greatly suppressed as shown in Figure 6B. The mechanism of such cleavage has been described in previous literatures24, and it is believed to go through an oxazolidine intermediate. PTA molecules can be sequenced by MS/MS and the fragmentation pattern is similar to that of peptides and peptoids, as shown in Figure 6C. PTA molecules synthesized with (S)-2-bromopropanoic acid-d4 generally give broader peak on MS and MS/MS spectra due to the presence of incomplete deuteration products such as (S)-2-bromopropanoic acid-d3 (Figures 7A and 7B). This could be used as an indication of the presence of the R chiral center (inverted from S during amination) during sequencing procedure. We also found that PTA molecules have a tendency to form more sodiated adducts than peptoid/peptide, therefore low sodium water (such as dionized water) and plastic apparatus are preferred (Figure 7C). Another by-product that could be observed in PTA synthesis is the acrylamide formed from the elimination of bromide during amination (Figure 7C). Once the acrylamide is formed, the sequence is terminated. This can be solved by lowering the concentration of the primary amine to 1 M in order to reduce the basicity of the solution. We recommend performing the chloranil test after each acylation step and using mass spectroscopy to check the product after each amination step to ensure the quality of the library.
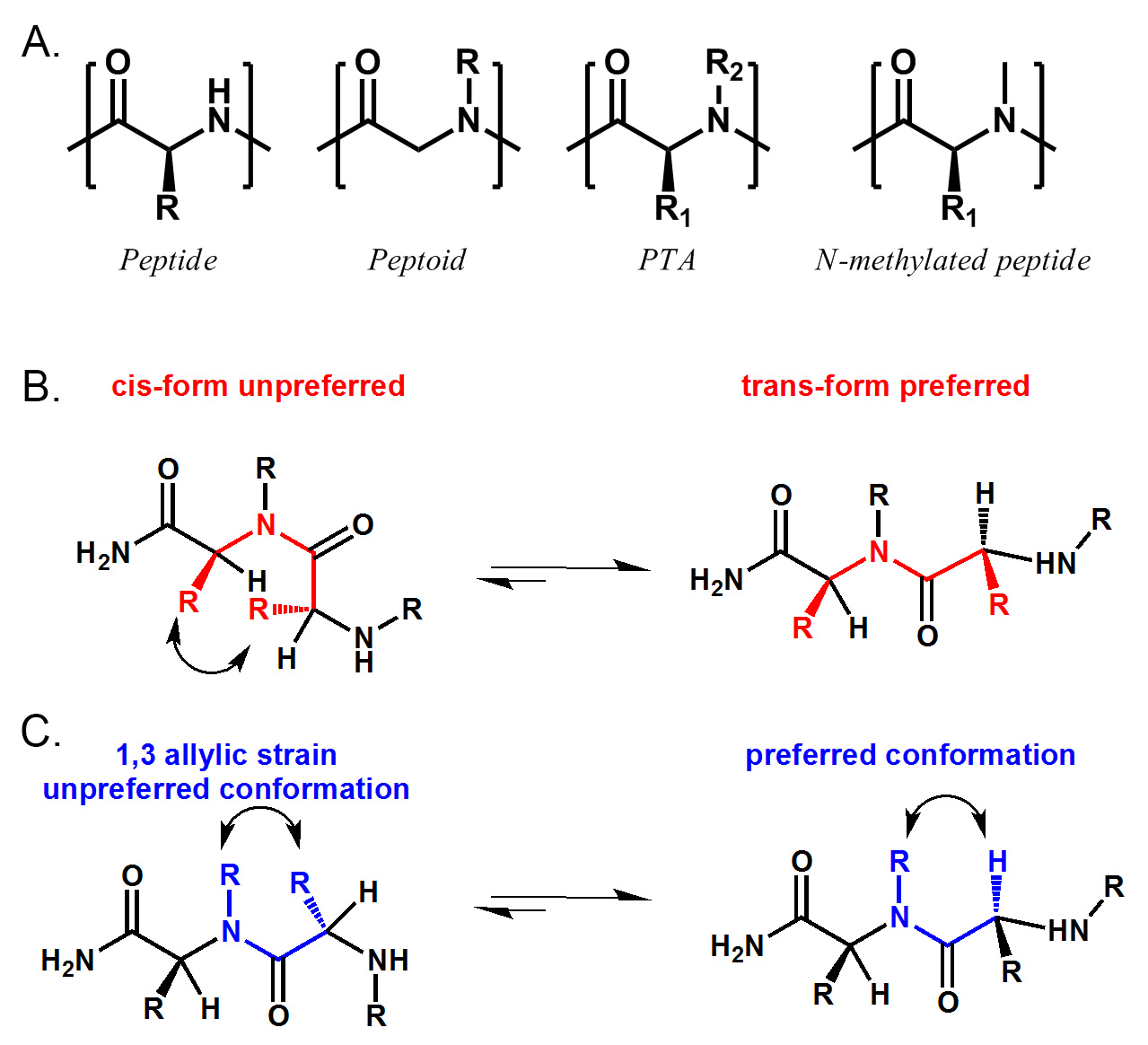
Figure 1. Structural comparison of peptide, peptoid, PTA and N-methylated peptide. PTA includes peptide (R2 = H), peptoid (R1 = H) and N-methylated peptide (R1 ≠ H, R2 = Me). B) PTA prefers trans amide bond conformation due to the sterical hindrance between two α-side chain. C) PTA also has a preferred conformation due to 1,3 allylic strain between N-substitute and α-side chain. Please click here to view a larger version of this figure.

Figure 2. Sub-monomer synthesis of peptoid (R1 = H) and PTA (R1 ≠ H). First step is the acid acylation of the amine. Second step is amination with primary amines. Please click here to view a larger version of this figure.

Figure 3. Reaction setup. A 250 ml three-neck round bottom flask is put in a dry ice/ethylene glycol bath. The middle neck is connected with a 150 ml pressure equalizing dropping funnel. The left and right necks are sealed with a flow control adapter and a septum with a long needle that allows argon flow pass through. Please click here to view a larger version of this figure.
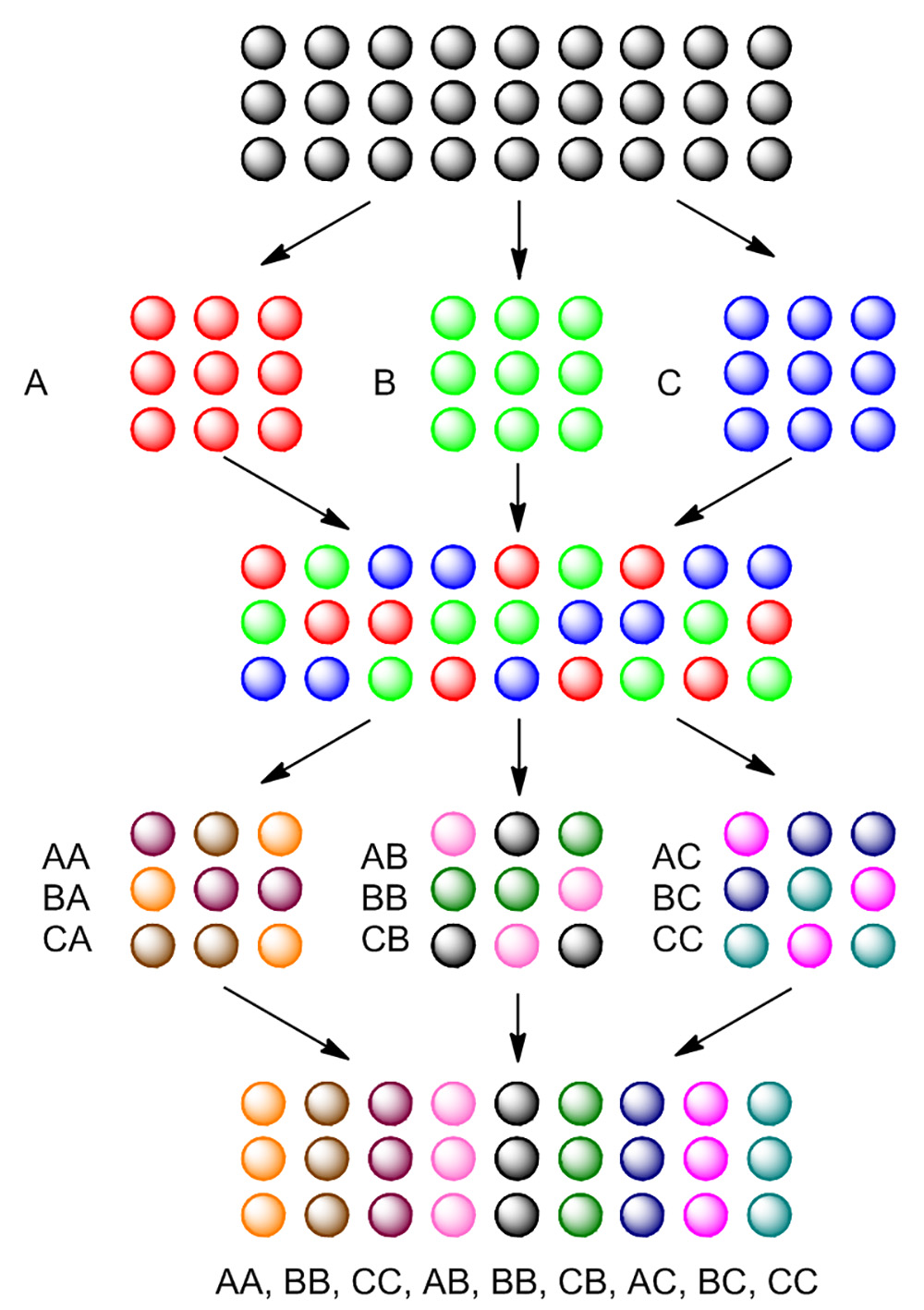
Figure 4. Basics of split-and-pool synthesis. Blank beads are split in three portions, treated separately with reactant A, B and C. After the first reaction, all three portions of beads are pooled together and mixed. Pooled beads are split again in three portions and again treated with same reactant for each individual portion. After the second reaction, 9 different compounds are synthesized. Please click here to view a larger version of this figure.

Figure 5. Library structure overview. Three PTAs are synthesized after the pentamer peptoid linker. Theoretical diversity, 33 X 73 = 9,261. Please click here to view a larger version of this figure.
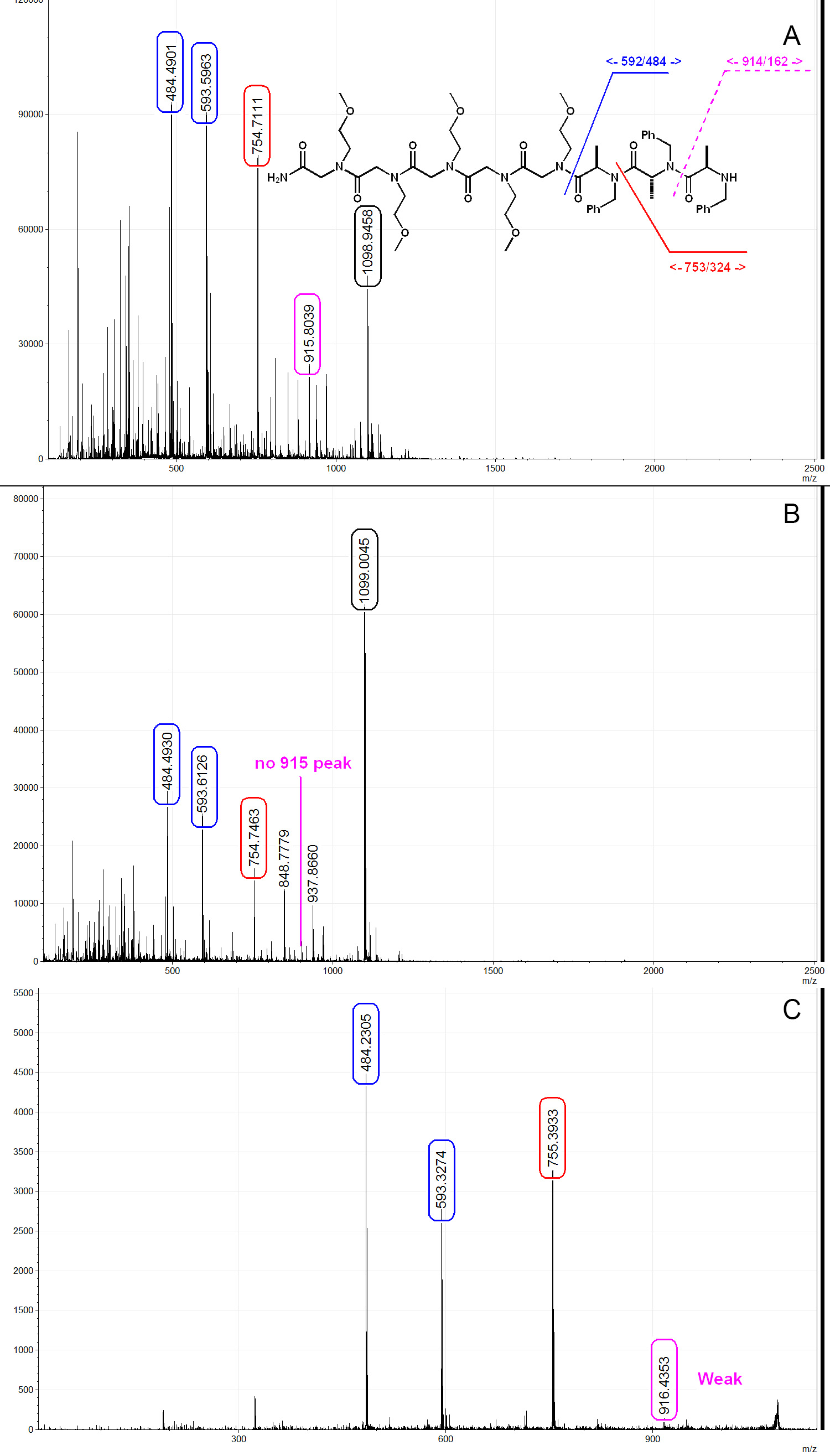
Figure 6. Typical MALDI mass spectra of a PTA trimer. A) Trimer PTA cleaved by 50% TFA/DCM under room temperature. PTA structure as shown, [M+1]+=1,077, [M+Na]+=1,098.9, PTA fragmentation from TFA cleavage can be clearly seen on the spectrum. B) Trimer cleaved by 50% TFA/DCM under optimized condition as described in the paper. TFA-induce acid degradation is greatly suppressed. C) MS/MS spectrum of the PTA trimer. Weak y7 (916) signal is observed, this is a typical fragmentation behavior for PTAs. Spectra analyzed and generated by mMass32. Please click here to view a larger version of this figure.

Figure 7. MALDI spectra of PTA synthesis with isotopic labeled monomer and typical by-products. A) MS spectra comparison of PTA molecules synthesized by blue: (R)-2-bromopropanoic [M+1]+=760 [M+Na]+=787 [M+K]+=803 and red: (S)-2-bromopropanoic acid-d4 [M+1]+=764 [M+Na]+=783 [M+K]+=799. B) MS/MS fragmentation patterns of the two molecules shown in A). Note that due to the presence of d1, d2 and d3 spices (incomplete deuteration of alanine), molecules synthesized by (S)-2-bromopropanoic acid-d4 generally give wider peaks. C) Red: Spectrum of a PTA dimer synthesized and cleaved under optimized condition. Blue: PTA dimer synthesized with 2 M methoxyethylamine solution and cleaved in normal filtered water. Please click here to view a larger version of this figure.
Discussion
Peptide tertiary amides (PTAs) are a superfamily of peptidomimetic oligomers. Besides the well-studied peptides, peptoids and N-methylated peptides, a large portion of compounds within this family remains understudied, majorly due to lack of synthetic method to access general N-alkylated peptides. Here we describe an efficient method to synthesize PTAs with chiral building blocks derived from amino acids. Previously, we have reported to use a new sub-monomer route to synthesis libraries of PTA molecules23. We have shown that PTAs are highly structured oligomers which possess conformational restrains through the backbone. When tested in vivo, PTA molecules showed improved cell permeability and therefore improved activity25. However, alongside with all the advantages, PTAs also come with some synthetic challenges, majorly from the acylation of secondary amines at hindered positions. The α-side chain which provides conformational restriction also brings steric hindrance for the following coupling step. In order to overcome these synthetic challenges, we performed an extensive optimization study and determined BTC as the best coupling reagent for this reaction23.
The key step of the synthetic route is BTC facilitated acylation of the secondary amine. During this process, BTC enables the generation of an acid chloride in situ26,27. Most other coupling reagents, which form either active esters or acid anhydrides as intermediates failed to provide clean acylation for continuous PTA synthesis. The existence of previous PTA units greatly impairs the coupling efficiency of the following PTA unit due to steric hindrance. Therefore, for the synthesis of multiple PTAs, a highly active intermediate with a small leaving group is strongly preferred. Among all the coupling conditions that we tested, in situ generated acid chloride by BTC works the best in our hand. However, even with the highly active acid chlorides, we recommend to avoid highly sterical hindered amines such as α-branched primary amines in library synthesis unless tested in advance. Aromatic amines such as anile often lead to incomplete substitution and thus should also be avoided. During the BTC coupling step, the solution should always be a pale yellow to orange color; a darker colored solution is an indication of overheating and may result in lower yield and increased formation of by-products. This can generally be solved by further cooling of the BTC solution, reduce the size of the reaction and faster transfer of the activated BTC/acid solution. Besides BTC, N-Ethoxycarbonyl-2-ethoxy-1,2-dihydroquinoline (EEDQ) is another coupling reagent that works well in PTA synthesis. The key intermediate is a mixed carbonic anhydride with a relatively small leaving group. In the case of EEDQ, 3 equivalent of EEDQ is dissolved together with the acid in DCM and then applied to the beads at room temperature. The reaction is normally done within 2 hr with mild shaking. This reaction releases CO2 during the reaction; therefore the reaction system should not be closed.
Another key step is the cleavage and the characterization of PTA molecules. A distinctive MS/MS fragmentation pattern has been observed when sequencing PTA molecules via MALDI-MS/MS (Figure 6). It consists with a low intensity of the last y ion (shown in purple in Figure 6C and increased intensity of y6, y5, y4, b2, b3 ions). Analogous patterns have been observed from N-methylated peptide fragmentation in the previous report28. Due to the increased stability of the intermediate oxazolidine, N-methylated peptides tend to give strong b ions28. Moreover, it is well known that N-methylated peptide are acid labile during both TFA cleavage and MALDI mass spectroscopy24,28,29. Mechanistic study has shown that due to the conformational restriction on the backbone, the carbonyl oxygen atom of the preceding residue is often in proximity of the carbonyl group at the cleavage site, thus promoting the formation of the oxazolidine intermediate29. For the reasons mentioned above, both PTAs and N-methylated peptides need to be cleaved from solid supports at low temperature using controlled concentrations of TFA26,30. In our experience, the most convenient cleavage method for individual PTA residues occurs at -20 °C with a pre-cooled, -20 °C solution of 50% TFA/DCM. This procedure greatly suppresses the formation of acid degraded products.
After mastering this technique, library with PTA monomer derived from other natural amino acids such as leucine, phenylalanine, glutamine, etc. can be synthesized as well. A high quality PTA library can be screened against various protein targets using our previously published on-bead screening protocols31. Hit compounds identified from the screening can be characterized by mass spectroscopy and resynthesized for further test using the protocol described above.
Disclosures
The authors have nothing to disclose.
Acknowledgements
The authors would like to thank Dr. Jumpei Morimoto and Dr. Todd Doran for valuable assistance. This work was supported by a contract from the NHLBI (NO1-HV-00242).
Materials
| 2,4,6 trimethylpyridine | ACROS | 161950010 | CAS:108-75-8 |
| 2-morpholinoethanamine | Sigma-Aldrich | 06680 | CAS:2038-03-1 |
| 48% HBr Water solution | ALFA AESAR | AA14036AT | CAS:10035-10-6 |
| Acetaldehyde | Sigma-Aldrich | 402788 | CAS:75-07-0 |
| Acetonitrile | Fisher | SR015AA-19PS | CAS:75-05-8 |
| Anhydrous Tetrahydrofuran (THF) | EMD | EM-TX0277-6 | CAS:109-99-9 |
| Benzylamine | Sigma-Aldrich | 185701 | CAS:100-46-9 |
| bis(trichloromethyl) carbonate (BTC) | ACROS | 258950050 | CAS:32315-10-9 |
| Bromoacetic acid | ACROS | 106570010 | CAS:79-08-3 |
| Chloranil | Sigma-Aldrich | 23290 | CAS:118-75-2 |
| Cyclohexanemethylamine | Sigma-Aldrich | 101842 | CAS:3218-02-8 |
| D2O | Cambridge Isotope | DLM-4-99.8-1000 | CAS:7789-20-0 |
| D-alanine | Anaspec | 61387-100 | CAS:338-69-2 |
| Dichloromethane (DCM) | Fisher | BJ-NS300-20 | CAS:75-09-2 |
| Dimethylformamide (DMF) | Fisher | BJ-076-4 | CAS:68-12-2 |
| Ethylene glycol | Oakwood | 44710 | CAS:107-21-1 |
| Isopentylamine | Sigma-Aldrich | W321907 | CAS:107-85-7 |
| KBr | ACROS | 424070025 | CAS:7758-02-3 |
| L-alanine | Anaspec | 61385-100 | CAS:56-41-7 |
| 3-Methoxypropylamine | Sigma-Aldrich | M25007 | CAS:5332-73-0 |
| 2-Methoxyethylamine | Sigma-Aldrich | 143693 | CAS:109-85-3 |
| N-(3-Aminopropyl)-2-pyrrolidinone | Sigma-Aldrich | 136565 | CAS:7663-77-6 |
| N,N'-Diisopropylcarbodiimide (DIC) | ACROS | 115211000 | CAS:693-13-0 |
| N,N-Diisopropylethylamine (DIPEA) | Sigma-Aldrich | D125806 | CAS:7087-68-5 |
| NaNO2 | ACROS | 424340010 | CAS:7631-99-4 |
| NAOD 40% solution in water | ACROS | 200058-506 | CAS:7732-18-5 |
| Piperidine | ALFA AESAR | A12442-AE | CAS:110-89-4 |
| Piperonylamine | Sigma-Aldrich | P49503 | CAS:2620-50-0 |
| Propylamine | Sigma-Aldrich | 240958 | CAS:107-10-8 |
| Trifluoroacetic acid | Sigma-Aldrich | 299537 | CAS:76-05-1 |
| α-Cyano-4-hydroxycinnamic acid | Sigma-Aldrich | 39468 | CAS:28166-41-8 |
| α-ketoglutarate | ALFA AESAR | AAA10256-22 | CAS:328-50-7 |
| Tentagel Resin with RINK linker | Rapp-Polymere | S30023 | |
| Alanine transaminase | Roche | 10105589001 | AKA: Glutamate-Pyruvate Transaminase (GPT) |
| Incubator | New Brunswick Scientific | Innova44 | |
| NMR | Bruker | 400MHz | |
| MALDI mass spectrometer | Applied Biosystems | 4800 MALDI-TOF/TOF | |
| Lyophilizer | SP Scientific | VirTis benchtop K | |
| Syringe reactor | INTAVIS | Reaction Column | 3ml, 5ml, 10ml, 20ml |
| Vacuum manifold | Promega | A7231 | Vac-Man |
References
- Xiao, X., Yu, P., Lim, H. -. S., Sikder, D., Kodadek, T. Design and Synthesis of a Cell-Permeable Synthetic Transcription Factor Mimic. Journal of Combinatorial Chemistry. 9, 592-600 (2007).
- Miller, S. M., et al. Proteolytic Studies of Homologous Peptide and N-Substituted Glycine Peptoid Oligomers. Bioorganic & Medicinal Chemistry Letters. 4, 2657-2662 (1994).
- Grauer, A., Konig, B. Peptidomimetics – A Versatile Route to Biologically Active Compounds. European Journal of Organic Chemistry. 30, 5099-5111 (2009).
- Zuckermann, R. N., Kerr, J. M., Kent, S. B. H., Moos, W. H. Efficient method for the preparation of peptoids [oligo(N-substituted glycines)] by submonomer solid-phase synthesis. Journal of the American Chemical Society. 114, 10646-10647 (1992).
- Figliozzi, G. M., Goldsmith, R., Ng, S. C., Banville, S. C., Zuckermann, R. N. Synthesis of N-substituted glycine peptoid libraries. Methods in Enzymology. 267, 437-447 (1996).
- Seebach, D., et al. beta-peptides: Synthesis by Arndt-Eistert homologation with concomitant peptide coupling. Structure determination by NMR and CD spectroscopy and by X-ray crystallography. Helical secondary structure of a beta-hexapeptide in solution and its stability towards pepsin. Helv Chim Acta. 79, 913-941 (1996).
- Lam, K. S., et al. A New Type of Synthetic Peptide Library for Identifying Ligand-Binding Activity. Nature. 354, 82-84 (1991).
- Simon, R. J., et al. Peptoids – a Modular Approach to Drug Discovery. Proceedings of the National Academy of Sciences of the United States of America. 89, 9367-9371 (1992).
- Burkoth, T. S., et al. Toward the synthesis of artificial proteins: the discovery of an amphiphilic helical peptoid assembly. Chem Biol. 9, 647-654 (2002).
- Alluri, P. G., Reddy, M. M., Bachhawat-Sikder, K., Olivos, H. J., Kodadek, T. Isolation of protein ligands from large peptoid libraries. Journal of the American Chemical Society. 125, 13995-14004 (2003).
- Lim, H. S., Archer, C. T., Kodadek, T. Identification of a peptoid inhibitor of the proteasome 19S regulatory particle. Journal of the American Chemical Society. 129, 7750-7751 (2007).
- Wrenn, S. J., Weisinger, R. M., Halpin, D. R., Harbury, P. B. Synthetic ligands discovered by in vitro selection. Journal of the American Chemical Society. 129, 13137-13143 (2007).
- Aina, O. H., Marik, J., Liu, R. W., Lau, D. H., Lam, K. S. Identification of novel targeting peptides for human ovarian cancer cells using "one-bead one-compound" combinatorial libraries. Mol Cancer Ther. 4, 806-813 (2005).
- Udugamasooriya, D. G., Dineen, S. P., Brekken, R. A., Kodadek, T. A Peptoid “Antibody Surrogate” That Antagonizes VEGF Receptor 2 Activity. Journal of the American Chemical Society. 130, 5744-5752 (2008).
- Shah, N. H., et al. Oligo( N-aryl glycines): A New Twist on Structured Peptoids. Journal of the American Chemical Society. 130, 16622-16632 (2008).
- Chongsiriwatana, N. P., et al. Peptoids that mimic the structure, function, and mechanism of helical antimicrobial peptides. Proceedings of the National Academy of Sciences of the United States of America. 105, 2794-2799 (2008).
- Paul, B., et al. N-Naphthyl Peptoid Foldamers Exhibiting Atropisomerism. Organic Letters. 14, 926-929 (2012).
- Crapster, J. A., Guzei, I. A., Blackwell, H. E. A peptoid ribbon secondary structure. Angewandte Chemie. 52, 5079-5084 (2013).
- Gorske, B. C., Stringer, J. R., Bastian, B. L., Fowler, S. A., Blackwell, H. E. New strategies for the design of folded peptoids revealed by a survey of noncovalent interactions in model systems. J Am Chem Soc. 131, 16555-16567 (2009).
- Stringer, J. R., Crapster, J. A., Guzei, I. A., Blackwell, H. E. Extraordinarily robust polyproline type I peptoid helices generated via the incorporation of alpha-chiral aromatic N-1-naphthylethyl side chains. J Am Chem Soc. 133, 15559-15567 (2011).
- Huang, K., et al. A threaded loop conformation adopted by a family of peptoid nonamers. Journal of the American Chemical Society. 128, 1733-1738 (2006).
- Lee, J. H., Kim, H. S., Lim, H. S. Design and Facile Solid-Phase Synthesis of Conformationally Constrained Bicyclic Peptoids. Organic Letters. 13, 5012-5015 (2011).
- Gao, Y., Kodadek, T. Synthesis and Screening of Stereochemically Diverse Combinatorial Libraries of Peptide Tertiary Amides. Chem Biol. 20, 360-369 (2013).
- Urban, J., Vaisar, T., Shen, R., Lee, M. S. Lability of N-alkylated peptides towards TFA cleavage. Int J Pept Protein Res. 47, 182-189 (1996).
- Rzuczek, S. G., Gao, Y., Tang, Z., Thornton, C. A., Kodadek, T., Disney, M. D. Features of Modularly Assembled Compounds That Impart Bioactivity Against an RNA Target. ACS Chemical Biology. 8 (10), 2312-2321 (2013).
- Thern, B., Rudolph, J., Jung, G. Triphosgene as highly efficient reagent for the solid-phase coupling of N-alkylated amino acids—total synthesis of cyclosporin O. Tetrahedron Letters. 43, 5013-5016 (2002).
- Sleebs, M. M., Scanlon, D., Karas, J., Maharani, R., Hughes, A. B. Total Synthesis of the Antifungal Depsipeptide Petriellin A. J Org Chem. 76, 6686-6693 (2011).
- Vaisar, T., Urban, J. Gas-phase fragmentation of protonated mono-N-methylated peptides. Analogy with solution-phase acid-catalyzed hydrolysis. Journal of Mass Spectrometry. 33, 505-524 (1998).
- Creighton, C. J., Romoff, T. T., Bu, J. H., Goodman, M. Mechanistic studies of an unusual amide bond scission. Journal of the American Chemical Society. 121, 6786-6791 (1999).
- Sewald, N., Sewald, N. Efficient, racemization-free peptide coupling of N-alkyl amino acids by using amino acid chlorides generated in situ–total syntheses of the cyclopeptides cyclosporin O and omphalotin A. Angewandte Chemie (International ed. in English). 41, 4661-4663 (2002).
- Astle, J. M., et al. Seamless Bead to Microarray Screening: Rapid Identification of the Highest Affinity Protein Ligands from Large Combinatorial Libraries. Chem Biol. 17, 38-45 (2010).
- Strohalm, M., Kavan, D., Novak, P., Volny, M., Havlicek, V. mMass 3: a cross-platform software environment for precise analysis of mass spectrometric data. Anal Chem. 82, 4648-4651 (2010).

