Zinc-finger Nuclease Enhanced Gene Targeting in Human Embryonic Stem Cells
Summary
Reporter cell lines offer a means to visualize, track and isolate cells of interest from heterogeneous populations. However, gene-targeting using conventional homologous recombination in human embryonic stem cells is extremely inefficient. Herein, we describe targeting CNS midbrain specific transcription factor PITX3 locus with EGFP using zinc-finger nuclease enhanced homologous recombination.
Abstract
One major limitation with current human embryonic stem cell (ESC) differentiation protocols is the generation of heterogeneous cell populations. These cultures contain the cells of interest, but are also contaminated with undifferentiated ESCs, non-neural derivatives and other neuronal subtypes. This limits their use in in vitro and in vivo applications, such as in vitro modeling for drug discovery or cell replacement therapy. To help overcome this, reporter cell lines, which offer a means to visualize, track and isolate cells of interest, can be engineered. However, to achieve this in human embryonic stem cells via conventional homologous recombination is extremely inefficient. This protocol describes targeting of the Pituitary homeobox 3 (PITX3) locus in human embryonic stem cells using custom designed zinc-finger nucleases, which introduce site-specific double-strand DNA breaks, together with a PITX3–EGFP-specific DNA donor vector. Following the generation of the PITX3 reporter cell line, it can then be differentiated using published protocols for use in studies such as in vitro Parkinson’s disease modeling or cell replacement therapy.
Introduction
Current embryonic stem cell (ESC) differentiation protocols result in heterogeneous cell populations, particularly with respect to the derivation of specific neuronal phenotypes. Thus, although cultures contain the desired cellular phenotype, other neuronal, non-neuronal and un-differentiated cell types are often present1. These characteristics limit the application of ESC derived cell sources for use in cell replacement therapy and in vitro disease modeling.
Genetic reporter cell lines offer a means to visualize, track and isolate cells of interest, provided that the expression of the reporter protein (RP) reproduces endogenous expression. Use of promoters immediately upstream of a gene coding region can be used to derive crude genetic reporters but such constructs lack the precise regulatory elements that control endogenous gene expression. In contrast, homologous recombination offers the opportunity to ensure high-fidelity expression of the RP. In the past, targeting vectors designed for homologous recombination at specific loci of interest have been used to target RPs in mouse ESCs (mESCs) using electroporation as a means of DNA delivery1,2. However the generation of reporter cell lines via conventional homologous recombination is extremely inefficient for human ESCs (hESCs), and thus has only been documented in a handful of cases (reviewed in3). By using an engineered chimeric protein containing a fusion of a FokI nuclease with site-specific zinc-finger motifs, known collectively as zinc-finger nucleases (ZFNs), DNA double-strand breaks can be introduced at pre-determined genomic loci. When a DNA vector with homology to both sides of the DNA double strand break is added, the genomic site can be repaired by homologous recombination, allowing the incorporation of the DNA donor sequence. This technique has proved useful for genomic editing in both human primary cells4,5 and hESCs6,7. More recent work has utilized transcription activator-like effector nucleases (TALENs), transcription factors used by plant pathogens8, to aid in the design of site-specific nucleases9.
In the following protocol we demonstrate the generation of a hESC reporter cell line by electroporation of an EGFP containing homologous targeting vector6 together with ZFNs for the human Pituitary homeobox 3 (PITX3) locus. Following antibiotic selection for 2-3 weeks, hESCs with correctly integrated DNA can be manually picked, expanded and screened initially via PCR, and subsequently validated by Southern blotting.
Protocol
All procedures are carried out in a sterile laminar flow hood. All media and solutions are equilibrated to 37 °C unless otherwise specified.
1. Expansion of Human PSCs (hESCs and hiPSCs, WA-09 used in this Protocol) for Transfection
Note: It is not necessary to expand hESCs on Matrigel or Geltrex. hESCs expanded on mouse embryonic fibroblasts (MEFs) can be used for electroporation or nucleofection following a MEF removal step (see section 2.8).
- Prepare 1 x 100 mm dish and 1 x 75cm2 flask of DR4 MEFs at 2.0 x 104 cells per cm2 in MEF media (see Table 1) on dishes pre-coated with 0.1% w/v gelatin.
Note: CF1, BLK6 or MF1 MEFs can be used, however, it is recommended that antibiotic resistant MEF lines are used, e.g., DR410. - The next day, passage hESCs onto a 100 mm dish pre-plated with MEFs prepared in section 1.1. To do this aspirate hESC media from cultures dishes, wash hESCs with Ca2+/Mg2+-free Hanks’ balanced salt solution (HBSS), add Dispase (see Table 2 for appropriate volumes) and place in a humidified incubator at 37°C/5% CO2. Following 3-5 min incubation, aspirate Dispase and gently wash 1 – 2 times with HBSS to remove MEFs. Add hESC media (see Table 3) and using a cell scraper, or 5 ml pipette, gently scrape the colonies off the surface of the culture dish.
- Collect the colony fragments in suspension from the culture dish into a 15 ml centrifuge tube. Rinse culture dish with hESC media and transfer to the 15 ml tube. Be careful to not break up clumps into single cells or small fragments.
- Centrifuge for 5 min at 160 x g to pellet colony fragments.
- Aspirate supernatant. Tap bottom of 15 ml tube with finger to loosen colony fragment pellet. Gently resuspend colony fragments in hESC media according to split ratio (normally a split ratio between 1:4 and 1:8 is appropriate). Be extremely careful not to render clumps into single cells or small fragments as this will affect replating efficiency.
- Prior to replating hESC fragments onto MEFs wash 1x with HBSS. Slowly add an appropriate amount (see Table 2) of cell suspension containing hESC fragments in hESC media to dishes pre-plated with MEFs.
- Place culture dishes onto shelf in incubator and move slowly back and forth and side to side to evenly distribute colony fragments.
- Replace hESC media daily.
- To prepare conditioned media from MEFs (MEF-CM) wash the 75 cm2 flask pre-plated with MEFs 1 x with HBSS. Add hESC media. Following 24 hr, collect conditioned media and replace with fresh hESC media. Repeat daily for a maximum of 12 days. Store MEF-CM at 4 °C until needed.
2. Preparation of Human ESCs for Transfection
- Observe the growth of hESCs under a microscope daily. When colonies become 40-50% confluent, plate 3 x 100 mm dishes with 2.0 x 105 MEFs per cm2.
- When colonies become 60 – 70% confluent they are ready to be transfected. ‘Groom’ colonies by removing differentiated colonies or colony cores.
Note: As we believe higher transfection and targeting efficiencies occur when cells are in log phase growth, it is suggested that electroporation be conducted before hESCs reach ≥ 70% confluency. - Wash hESC colonies with HBSS and add Accutase.
Note: It is not necessary to pre-treat hESCs with ROCK inhibitor Y-27632 as sufficient inhibition will occur when incubated with Y-27632 during step 2.7 - Incubate for 10 – 20 min at 37°C/5% CO2 until colonies are rendered single cells as viewed under a microscope. Triturate cells with a 1 ml pipette.
- Add hESC media and filter into a 15 ml conical tube using a 40 μm cell strainer to remove cell clumps.
- Centrifuge at 160 x g for 5 min. Resuspend cell pellet in fresh hESC media and repeat to remove any residual Accutase solution.
- Resuspend cells in hESC media containing 10 μM Y-27632 and transfer the total cell suspension to a 100 mm dish pre-coated with gelatin.
- Incubate at 37 °C/5% CO2 for a minimum of 30 min.
- During this time MEFs will adhere to the gelatin coated dish. Collect non-adherent hESCs using a 1 ml pipette into a fresh 15 ml centrifuge tube and determine cell density using a hemocytometer.
3. Electroporation of hESCs
- Filter CM-MEF media using a 0.22 μm filter and add 10 ng/ml recombinant human fibroblast growth factor 2 (rhFGF2).
- Transfer 7.5 x 106 hESCs from section 2.9 to a new 15 ml conical tube and centrifuge at 160 x g for 5 min.
- During this time set up the electroporator: Select program for exponential electroporation and set the following parameters, Voltage: 250 V, Capacitance: 500 μF, and Resistance: ∞-.
- Aspirate the supernatant and re-suspend cell pellet in 0.8 ml of ice-cold 0.22 μm filtered HBSS. Transfer the solution to a sterile 0.4 cm electroporation cuvette.
- Add TV-hPITX3-forward targeting construct (30 μg DNA) and mix gently with a 200 μl pipette. Incubate on ice for 5 min. During this time remove one aliquot of PITX3 ZFN mRNA and defrost on ice. When defrosted transfer 7.5 μl of ZFN mRNA to DNA/cell mixture in electroporation cuvette. Mix gently with a 200 μl pipette.
Note: The locus being targeted is the main factor that contributes to targeting efficiency. Loci encoding silent genes, genes not expressed in ESCs, such as PITX3 are difficult to target using conventional homologous recombination6. Thus, this protocol utilizes custom designed ZFNs for the human PITX3 locus to induce a DNA double strand break. - Electroporate by following the on screen instructions on the electroporator.
- Using a 200 μl pipette transfer the contents from the electroporation cuvette to a 15 ml tube containing 10 ml hESC media. Wash the cuvette twice with 200 μl of hESC media each time and transfer to a 15 ml tube. There will be “mucous-like” debris consisting of DNA and protein from the lysed cells which will negatively impact cell survival. Centrifuge at 160 x g for 5 min to remove this debris.
- Aspirate the supernatant and gently tap cell pellet to loosen. Re-suspend in 30 ml supplemented MEF-CM containing 10 μM Y-27632 and replate onto 3 x 100 mm dishes preplated with MEFs.
- Change the media daily. On day 2, Y-27632 can be withdrawn and cells grown in hESC media only.
- After 2-3 days the hESCs should be 50-70% confluent. At this point begin antibiotic selection (e.g., 0.5 μg/ml puromycin).
- Maintain selection by adding fresh hESC/puromycin media daily. After ~7 days small colonies should be visible. If antibiotic resistant MEFs are not used, MEFs can be supplemented by adding an additional 1.0 x 104 cells per cm2.
4. Picking Transgenic hESC Clones
- Approximately 14 – 21 days following selection (when colonies have reached ~1 mm diameter) prepare plates for expansion and screening. Begin by defrosting an aliquot of Matrigel or Geltrex at 4°C for a minimum of 5 hr. Dilute as per manufacturers’ instructions and add 75 μl to each well of 3 x 96-well plates. Prepare 3 x 48-well plates by plating MEFs at 2.0 x 104 cells per cm2. Incubate both plates O/N at 37 °C/5% CO2.
- The following day dissect colonies into 16-25 pieces using a 21 G needle on the end of a 2 ml pipette by cutting a grid.
- Aspirate Matrigel from 96-well plates and replace with supplemented CM-MEF media. Wash MEFs in 48-well plates 1 x with HBSS and add hESC media.
- Using a 200 μl pipette gently transfer 1/3 of the dissected colony into a 48-well plate for expansion. Transfer the other 2/3 of the dissected colony into a corresponding 96-well plate containing supplemented CM-MEF media.
- Maintain both plates by changing the media daily (supplemented CM-MEF media for 96-well matrigel coated plates and hESC media for 48-well plates containing MEFs; both supplemented with 0.5 μg/ml puromycin).
- Expand until 96-well plates are 100% confluent and then initially screen via PCR.
5. Screening and Expansion of Positive Clones
- To prepare genomic DNA (method based on 11) remove all media from plates by inverting, then blot onto paper towels.
- Rinse each well twice with RT PBS (100 µl/well each wash), invert and blot as for step 5.1. Place plates at -80 °C for 1 – 2 hr to aid in cell lysis (plates can be stored indefinitely like this).
- While cells are at -80 °C make Sarcosyl Lysis Buffer (see Table 4) (50 µl/well). To lyse cells, warm plates for 5 min at RT then pipette lysis buffer into each well. Seal the edges of each plate with tape, place in a sealable container containing moist paper towels to create humidity, and incubate O/N at 60 °C.
- After incubation add 150 µl of 5M NaCl to 10 ml of -20 °C absolute ethanol, mix well, and add 100 µl to each well (Note: ethanol will go cloudy when NaCl is added). Tap plates gently to mix (do not pipette) and leave for a minimum of 30 min at RT. During this time the solution will go clear and the DNA will precipitate.
- Remove solution as in step 5.1 then add 150 µl of 70% ethanol to each well. Invert plate gently and blot as before. Repeat this step a further 2x.
Note: Because the precipitated DNA naturally adheres to the tissue culture plastic it does not become dislodged during these washing steps.) - After the final wash briefly centrifuge plates at top speed to concentrate the DNA to the bottom of each well then air-dry at RT (or 37 °C) for at least 30 min.
- Add 40 – 50 µl T0.1E (pH8.0) (see Table 5) or TE (pH8.0) to each well and place plates at 65 °C for 1 hr or at 4 °C O/N before using for PCR. Resuspend DNA by gently tapping the sides of the plate. Use 1-2 µl of re-suspended DNA/10 µl PCR reaction.
- Using the Expand Long Template PCR System with hPITX3 L arm gen. F and hPITX3 L arm GFP R primers perform initial screening of clones at 58 °C annealing, 45 sec extension, 35 cycles.
- Electrophorese samples with Hyperladder 1kb in a 1% agarose gel containing GelRed at 100 V for 45 min and detect positive clones via UV illumination.
- Validate PCR positive clones by digesting genomic DNA from expanded clones (see below) with HindIII and hybridizing probes 5’ and 3’ of the vector homology arms to a Southern blot (as previously described in 1). Probes are generated by PCR using primer pairs hPITX3 5’ probe F and R, and hPITX3 3’ probe F and R, digested with EcoRI and ligated into a general cloning vector.
- Depending on the number of positive clones, prepare 2 wells of a 6-well plate by pre-plating with MEFs (2.0 x 104 cells per cm2).
- The next day pretreat positive clones in the 48-well plate with 10 μM Y-27632 for a minimum of 1 hr.
- Wash positive clones with HBSS and add 300 μl of Accutase.
- When colonies are rendered single cells, add 1 ml of hESC media and collect into a 15 ml tube.
- Change the media daily. On day 2, Y-27632 can be withdrawn and cells grown in hESC media.
- When colonies are 70 – 80% confluent, subculture and freeze the remainder of the colony fragments.
Representative Results
Following co-electroporation of the custom designed PITX3 zinc-finger pair together with the PITX3–EGFP-specific DNA donor vector, and subsequent puromycin selection, positive hESC clones were initially detected via genomic PCR screening (Figure 1). Southern blot hybridization of these PCR positive clones with 5’ and 3’ specific probes confirmed correct targeting to exon 1 of the PITX3 locus (Figure 2), with an efficiency of 19%. Immunofluorescent images shown in Figure 3 demonstrate EGFP expression driven by the PITX3 promoter following differentiation using a published PA6 stromal cell differentiation protocol12,13. Co-localization of EGFP could be observed with markers of midbrain dopaminergic neurons such as FOXA2 (red) (Figure 3A) and tyrosine hydroxylase (TH; red) (Figure 3B), while importantly no co-localization could be observed with the GABAergic marker, gamma-aminobutyric acid (GABA; red) (Figure 3C).
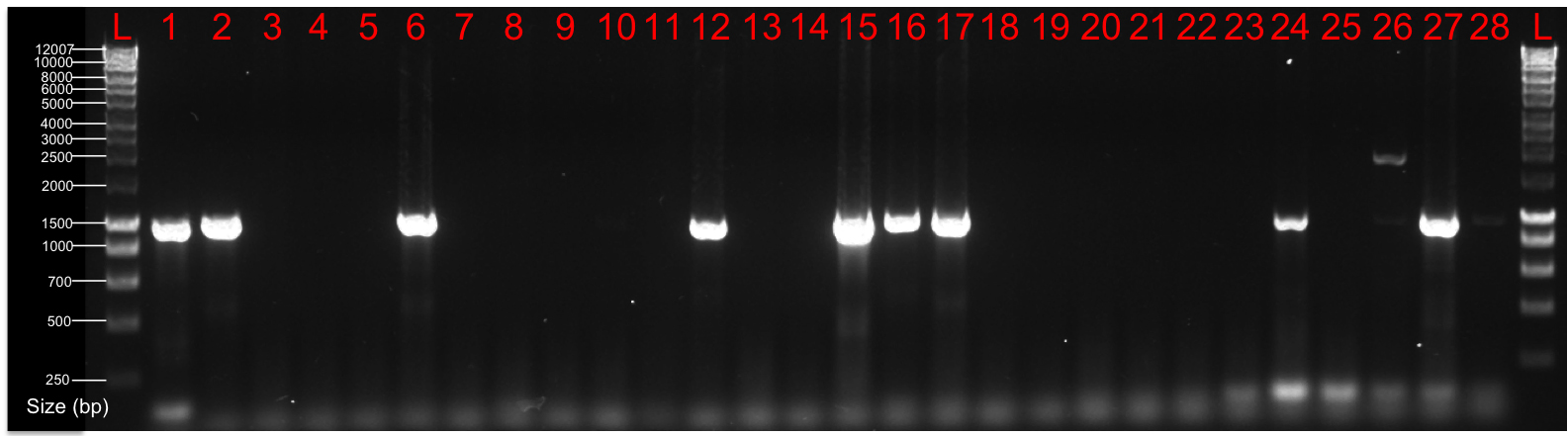
Figure 1. Preliminary genomic PCR screening. Following colony picking and expansion, preliminary genomic PCR screening detected numerous positive clones as shown by the PCR product (band at 984 bp). (Clones were screened using primers hPITX3 L arm gen. F, which lies just outside the left targeting arm, and hPITX3 L arm GFP R which is located within the EGFP gene.) Right-hand and left-hand side lanes are a 1 kb ladder. Please click here to view a larger version of this figure.

Figure 2. Validation of EGFP targeting to the hPITX3 locus using zinc finger nucleases in hESCs. (A) Schematic of the Southern blot strategy employed to validate targeting. The top panel schematic illustrates the native hPITX3 locus before targeting. The bottom panel schematic depicts the hPITX3 locus following correct insertion of the EGFP transgene. The red lines under each panel illustrate the sites at which the 5’ and 3’ Southern blot probes will bind, and correspond to the bands in Figure 2B. (B) Representative Southern blot of correctly targeted colonies selected from PCR pre-screening. Abbreviations: EGFP, enhanced green fluorescent protein; PGK, human phosphoglycerol kinase promoter; PURO, puromycin resistance gene. Other features: loxP sites, purple; polyadenylation sequence, blue, hPITX3 exons, orange. Please click here to view a larger version of this figure.
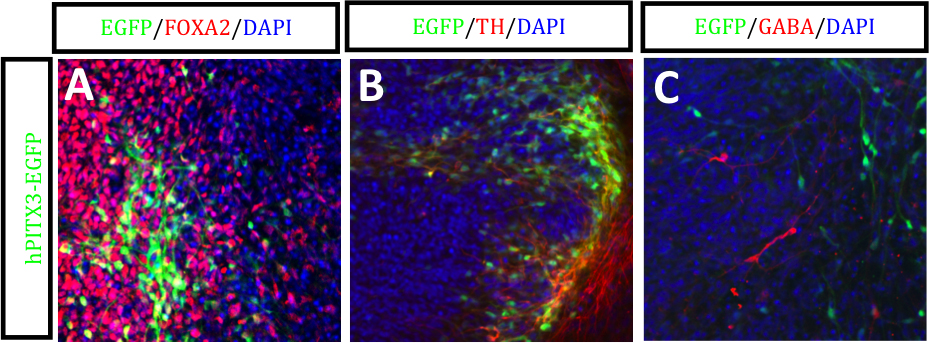
Figure 3. hPITX3-EGFP hESCs differentiated under PA6/LSB/SAG/FGF8 conditions. Immunofluorescent images of EGFP labeled with (A) FOXA2 (red), (B) TH (red) and (C) GABA (red). Counterstained with DAPI (blue). Scales bars, 50 μm. Please click here to view a larger version of this figure.
Following co-electroporation of the custom designed PITX3 zinc-finger pair together with the PITX3–EGFP-specific DNA donor vector, and subsequent puromycin selection, positive hESC clones were initially detected via genomic PCR screening (Figure 1). Southern blot hybridization of these PCR positive clones with 5’ and 3’ specific probes confirmed correct targeting to exon 1 of the PITX3 locus (Figure 2), with an efficiency of 19%. Immunofluorescent images shown in Figure 3 demonstrate EGFP expression driven by the PITX3 promoter following differentiation using a published PA6 stromal cell differentiation protocol12,13. Co-localization of EGFP could be observed with markers of midbrain dopaminergic neurons such as FOXA2 (red) (Figure 3A) and tyrosine hydroxylase (TH; red) (Figure 3B), while importantly no co-localization could be observed with the GABAergic marker, gamma-aminobutyric acid (GABA; red)(Figure 3C).

Figure 1. Preliminary genomic PCR screening. Following colony picking and expansion, preliminary genomic PCR screening detected numerous positive clones as shown by the PCR product (band at 984 bp). (Clones were screened using primers hPITX3 L arm gen. F, which lies just outside the left targeting arm, and hPITX3 L arm GFP R which is located within the EGFP gene.) Right-hand and left-hand side lanes are a 1 kb ladder. Please click here to view a larger version of this figure.

Figure 2. Validation of EGFP targeting to the hPITX3 locus using zinc finger nucleases in hESCs. (A) Schematic of the Southern blot strategy employed to validate targeting. The top panel schematic illustrates the native hPITX3 locus before targeting. The bottom panel schematic depicts the hPITX3 locus following correct insertion of the EGFP transgene. The red lines under each panel illustrate the sites at which the 5’ and 3’ Southern blot probes will bind, and correspond to the bands in Figure 2B. (B) Representative Southern blot of correctly targeted colonies selected from PCR pre-screening. Abbreviations: EGFP, enhanced green fluorescent protein; PGK, human phosphoglycerol kinase promoter; PURO, puromycin resistance gene. Other features: loxP sites, purple; polyadenylation sequence, blue, hPITX3 exons, orange. Please click here to view a larger version of this figure.

Figure 3. hPITX3-EGFP hESCs differentiated under PA6/LSB/SAG/FGF8 conditions. Immunofluorescent images of EGFP labeled with (A) FOXA2 (red), (B) TH (red) and (C) GABA (red). Counterstained with DAPI (blue). Scales bars, 50 μm. Please click here to view a larger version of this figure.
| Substance | ml/100 ml | Catalogue No. (Supplier) |
| DMEM | 89 | 10566-016 (Life Technologies) |
| Fetal Bovine Serum | 10 | 16140-071 (Life Technologies) |
| Pen/Strep | 1 | 15070-063 (Life Technologies) |
Table 1.
| Component | 100 mm dish | 60 mm dish | 6-well | 48-well | 96-well | 75cm2 flask |
| HBSS | 5 ml | 2 ml | 1 ml | 0.5 ml | 0.1 ml | 8 ml |
| Accutase | 5 ml | 2 ml | 1 ml | 0.5 ml | 0.1 ml | 8 ml |
| Dispase | 5 ml | 2 ml | 1 ml | – | – | – |
| hESC media | 10 ml | 6 ml | 3 ml | 0.5 ml | 0.2 ml | 15 ml |
| Matrigel | – | – | – | – | 0.1ml | – |
Table 2.
| Substance | ml/100 ml | Catalogue No. (Supplier) |
| DMEM/F12 | 80 | 11320-033 (Life Technologies) |
| KSR | 20 | 10828-028 (Life Technologies) |
| NEAA | 1 | 11140-050 (Life Technologies) |
| GlutaMAX | 1 | 35050-061 (Life Technologies) |
| Pen/Strep | 1 | 15070-063 (Life Technologies) |
| ß-mercaptoethanol | 0.1 | 21985-023 (Sigma-Aldrich) |
| rhFGF2 | 7 ng/ml [final] | 233-FB-025/CF (R&D Systems) |
Table 3.
| Substance | Concentration | Catalogue No. (Supplier) |
| N-Lauroylsarcosine sodium salt solution (Sarcosyl) | 0.5% (w/v) | 61747 (Sigma-Aldrich) |
| EDTA | 10.0 mM | E9884 (Sigma-Aldrich) |
| NaCl | 10.0 mM | S3014 (Sigma-Aldrich) |
| Tris-Cl (pH7.5) | 10.0 mM | T3253 (Sigma-Aldrich) |
| Proteinase K | 1.0 mg/ml | BIO-37084 (Bioline Australia) |
Table 4.
| Substance | Concentration | Catalogue No. (Supplier) |
| Tris-Cl (pH8.0) | 10.0 mM | T3253 (Sigma-Aldrich) |
| EDTA (pH8.0) | 0.1 mM | E9884 (Sigma-Aldrich) |
Table 5.
Discussion
The generation of reporter cell lines offers a powerful means to track, visualize and isolate cells of interest from a heterogeneous population derived from hESCs. However, gene targeting via conventional homologous recombination has proven to be extremely inefficient for human ESCs3. In this protocol we describe a relatively simple technique for introducing a RP into exon one of the PITX3 locus, in hESCs using a widely available targeting vector together with ZFNs. In our hands the gene targeting system is efficient, with 19% of puromycin-resistant clones containing a correctly targeted PITX3 locus (similar to that previously published6).
A limitation with using ZFNs to facilitate gene targeting is the difficulty in designing ZFNs from scratch in house. Thus, in this protocol we used custom designed commercially available ZFNs, which can be expensive to acquire. More recently, the TALEN system has been used for gene targeting using homologous recombination9. The use of TALENs greatly reduces the cost of gene editing and can be designed in-house, expediting the process. Although not explicitly demonstrated in this study, this protocol can be easily modified to incorporate the use of TALENs as the methodology is the same, except TALEN plasmids are used in Step 3.5 instead of ZFN mRNA, as shown in 9.
When using this technique it is imperative to ensure that hPSCs are in log phase growth, (e.g., cultures are not 70% or more confluent). We believe that this is critical as the plasmid gains access to the nucleus following nuclear envelope breakdown during cell division. Moreover, the DNA donor vector will only incorporate into DNA following DNA synthesis as this process exploits homology-directed repair during division to incorporate the donor DNA into the genome.
Another potential limitation with the ZFN approach is the introduction of DNA breaks elsewhere in the genome. Although Southern hybridization with external 5’ and 3’ probes detects correct integration it will not detect additional integration events or error-prone repair elsewhere in the genome. Extra integration events can be examined by Southern hybridization using vector-specific probes (e.g., EGFP) whereas off-target cleavage of the ZFN pairs can be detected using SELEX3,6.
Although we did not observe any obvious effects in the development of midbrain dopaminergic neurons that could be directly attributed to the inactivation of one PITX3 gene, researchers should bear in mind that any targeting event where one copy of a gene is disrupted may be detrimental to the development of that cell and should take this into account when analyzing results. Whether this is the case will depend largely on the gene being targeted and may only be ascertained by performing the experiment.
Using a PA6 based differentiation protocol12,13 we were able to confirm the fidelity of the reporter system by immunofluorescence (Figure 3) and qPCR analysis of EGFP positive vs. negative populations following fluorescence-activated cell sorting (data not shown). As PITX3 is a transcription factor specific to midbrain dopaminergic neurons14, the engineering of a human PITX3 reporter cell line will facilitate the use of PITX3+ midbrain dopaminergic neurons derived from hESCs in studies of either in vitro PD modeling or cell replacement therapy.
Disclosures
The authors have nothing to disclose.
Acknowledgements
Work described in this protocol was made possible by funding from the Victorian Government as part of an collaborative alliance with the Californian Institute for Regenerative Medicine (CIRM).
Materials
| Mouse Embryonic Fibroblasts (DR4) | GlobalStem | GSC-6004G | |
| Gelatin | Sigma | G1393-100ML | |
| HBSS, no Calcium, no Magnesium, no Phenol Red (HBSS) | Life Technologies | 14175-095 | |
| Dispase | StemCell Technologies | 7923 | |
| Cell Scraper | Corning | 3008 | |
| Accutase | Life Technologies | A11105-01 | |
| Y27632 | Axon Medchem | Axon 1683 | |
| Cell Strainer – 40um | BD Biosciences | 352340 | |
| 33mm – 0.22 um filter | Millipore | SLGP033RS | |
| rhFGF2 | R&D Systems | 233-FB-025/CF | |
| Electroporator | BioRad | 165-2661 | |
| 0.4cm electroporation cuvette | BioRad | 165-2088 | |
| Human TV-hPITX3-forward targeting construct | Addgene | 31942 | |
| ZFN Kit; Human PITX3 | Sigma-Aldrich | CKOZFN1050-1KT | |
| Puromycin | Invivogen | ant-pr-1 | |
| Matrigel | BD Biosciences | 354277 | |
| Geltrex | Life Technologies | A1569601 | |
| NaCl | Sigma-Aldrich | S3014 | |
| Dulbecco's Modified Eagle Medium (DMEM) | Life Technologies | 10566-016 | |
| Fetal Bovine Serum, Qualified, Heat Inactivated, US Origin (FBS) | Life Technologies | 16140-071 | |
| Pen/Strep | Life Technologies | 15070-063 | |
| Dulbecco's Modified Eagle Medium/Nutrient Mixture F-12 (DMEM/F12) | Life Technologies | 11320-033 | |
| KnockOut Serum Replacement (KSR) | Life Technologies | 10828-028 | |
| MEM Non-Essential Amino Acids (NEAA) | Life Technologies | 11140-050 | |
| GlutaMAX | Life Technologies | 35050-061 | |
| ß-mercaptoethanol | Life Technologies | 21985-023 | |
| N-Lauroylsarcosine sodium salt solution (Sarcosyl) (30%) | Sigma-Aldrich | 61747 | |
| EDTA | Sigma-Aldrich | E9884 | |
| Trizma Hydrochloride | Sigma-Aldrich | T3253 | |
| Proteinase K solution (20mg/ml) | Bioline Australia | BIO-37084 | |
| Expand Long Template PCR System | Roche Australia | 11681834001 | |
| hPITX3 L arm gen. F (primer): 5’ TGTCCTAAGGAGAATGGGTAACAGACA 3’ | GeneWorks, Australia | ||
| hPITX3 L arm GFP R (primer): 5’ ACACGCTGAACTTGTGGCCGTTTA 3’ | GeneWorks, Australia | ||
| HyperLadder 1kb | Bioline Australia | BIO-33025 | |
| GelRed | Jomar Bioscience, Australia | 41003 |
References
- Nefzger, C. M., et al. Lmx1a Allows Context-Specific Isolation of Progenitors of GABAergic or Dopaminergic Neurons During Neural Differentiation of Embryonic Stem Cells. Stem Cells. 30 (7), 1349-1361 (2012).
- Zeng, W. R., Fabb, S. R. A., Haynes, J. M., Pouton, C. W. Extended periods of neural induction and propagation of embryonic stem cell-derived neural progenitors with EGF and FGF2 enhances Lmx1a expression and neurogenic potential. Neurochemistry International. 59 (3), 394-403 (2011).
- Collin, J., Lako, M. Concise review: putting a finger on stem cell biology: zinc finger nuclease-driven targeted genetic editing in human embryonic stem cells. Stem Cells. 29 (7), 1021-1033 (2011).
- Carroll, D. Progress and prospects: zinc-finger nucleases as gene therapy agents. Gene Therapy. 15 (22), 1463-1468 (2008).
- Urnov, F. D., et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature. 435 (7042), 646-651 (2005).
- Hockemeyer, D., et al. Efficient targeting of expressed and silent genes in human ESCs and iESCs using zinc-finger nucleases. Nature Biotechnology. 27 (9), 851-857 (2009).
- Lombardo, A., et al. Gene editing in human stem cells using zinc finger nucleases and integrase-defective lentiviral vector delivery. Nature Biotechnology. 25 (11), (2007).
- Boch, J., Bonas, U. Xanthomonas AvrBs3 family-type III effectors: discovery and function. Annual Review of Phytopathology. 48, 419-436 (2010).
- Hockemeyer, D., et al. Genetic engineering of human embryonic cells using TALE nucleases. Nature Biotechnology. 29 (8), 731-734 (2011).
- Tucker, K. L., Wang, Y., Dausman, J., Jaenisch, R. A transgenic mouse strain expressing four drug-selectable marker genes. Nucleic Acids Research. 25 (18), 3745-3746 (1997).
- Ramírez-Solis, R., Rivera-Pérez, J., Wallace, J. D., Wims, M., Zheng, H., Bradley, A. Genomic DNA microextraction: a method to screen numerous samples. Analytical Biochemistry. 201 (2), 331-335 (1992).
- Kawasaki, H., et al. Induction of midbrain dopaminergic neurons from ES cells by stromal cell-derived inducing activity. Neuron. 28 (1), 31-40 (2000).
- Denham, M., Thompson, L. H., Leung, J., Pébay, A., Björklund, A., Dottori, M. Gli1 is an inducing factor in generating floor plate progenitor cells from human embryonic stem cells. Stem Cells. 28 (10), 1805-1815 (2010).
- Smidt, M. P., et al. A homeodomain gene Ptx3 has highly restricted brain expression in mesencephalic dopaminergic neurons. Proceedings of the National Academy of Sciences U.S.A. 94, 13305-13310 (1997).

