Immunofluorescence Analysis of Endogenous and Exogenous Centromere-kinetochore Proteins
Summary
Here we report protocols to detect endogenous and exogenous centromere-kinetochore proteins in human cells and quantify these protein levels at centromeres-kinetochores by indirect immunofluorescent staining through the use of fixation (paraformaldehyde, acetone, or methanol fixation).
Abstract
“Centromeres” and “kinetochores” refer to the site where chromosomes associate with the spindle during cell division. Direct visualization of centromere-kinetochore proteins during the cell cycle remains a fundamental tool in investigating the mechanism(s) of these proteins. Advanced imaging methods in fluorescence microscopy provide remarkable resolution of centromere-kinetochore components and allow direct observation of specific molecular components of the centromeres and kinetochores. In addition, methods of indirect immunofluorescent (IIF) staining using specific antibodies are crucial to these observations. However, despite numerous reports about IIF protocols, few discussed in detail problems of specific centromere-kinetochore proteins.1-4 Here we report optimized protocols to stain endogenous centromere-kinetochore proteins in human cells by using paraformaldehyde fixation and IIF staining. Furthermore, we report protocols to detect Flag-tagged exogenous CENP-A proteins in human cells subjected to acetone or methanol fixation. These methods are useful in detecting and quantifying endogenous centromere-kinetochore proteins and Flag-tagged CENP-A proteins, including those in human cells.
Introduction
"Centromeres" were classically defined as regions of suppressed meiotic recombination in genetics and later recognized as the primary constriction of mitotic chromosomes, which plays an essential role in accurate chromosome segregation during mitosis. "Kinetochores" were described as the multilayered structures that bind to microtubules at the surface of centromeres, as revealed by electron microscopy; "kinetochores" were later defined as the macromolecular complex that localizes at the centromere of mitotic chromosomes. Despite a dramatic divergence of centromeric DNA sequences among vertebrates, kinetochore structure and composition are highly conserved. A dynamic interaction between spindle microtubules and the kinetochore is required for faithful segregation of chromosomes during mitosis, and defects in centromere-kinetochore function lead to aneuploidy and thereby cancer.
The centromere in most eukaryotes has no defined DNA sequence, but is composed of large arrays (0.3-5 Mb) of repetitive alphoid DNA consisting of 171-bp α-satellite DNA. Except in budding yeast, centromere identity is achieved not by the DNA sequence but by the presence of a special nucleosome that contains the histone H3 variant CenH3 (CENtromere Protein A [CENP-A] in humans).5 CENP-A nucleosomes localize to the inner plate of mammalian kinetochores7 and bind to the 171-bp α-satellite DNA. Active centromeres require CENP-A-containing nucleosomes to direct the recruitment of a constitutive centromere-associated network (CCAN) and the kinetochore proteins, which together regulate the attachment of chromosomes to the mitotic spindle and subsequent cycle progression through the spindle checkpoint.
In light of the above evidence, CENP-A has been proposed to be the epigenetic mark of the centromere8; however, the process by which CENP-A is incorporated into centromeric DNA and the factors responsible for this incorporation have not yet been well characterized. A short centromere-targeting domain (CATD) resides in the histone fold region of CENP-A, and replacement of the corresponding region of H3 with the CATD is sufficient to direct H3 to the centromere.9 Several studies suggested functional roles for post-translational modification (PTM) of CENP-A12-16; however, the molecular mechanisms of these PTMs of CENP-A in the recruitment to centromeres have not yet been elucidated. We previously reported that CUL4A-RBX1-COPS8 E3 ligase activity is required for CENP-A K124 ubiquitylation and localization of CENP-A to centromeres.17
The discovery and characterization of kinetochore proteins have led to new insight regarding chromosome segregation.18 More than 100 kinetochore components have been identified in vertebrate cells by various approaches.19,20 An understanding of how kinetochores assemble and function also comes from the characterization of the cellular functions of each centromere-kinetochore proteins and protein-protein network within cells.19 Direct visualization and advanced imaging methods in fluorescence microscopy provide remarkable resolution of centromere-kinetochore components and allow direct observation of specific molecular components of the centromeres and kinetochores. In addition, methods of indirect immunofluorescent (IIF) staining using specific antibodies are crucial to these observations. However, despite numerous reports about IIF protocols, few discussed in detail problems of specific centromere-kinetochore proteins.1-4 Thus, developing and reporting methods of IIF staining and a quantitative IIF assay to specifically analyze each centromere-kinetochore protein is extremely important. In IIF staining, one should proceed with the staining protocol to avoid loss of the protein of interest or the rest of the cell. However, fixation destroys antigenic sites occasionally, and different antibody-antigen combinations work poorly with one fixative, but very well with another,21 and choice of fixative depends largely on the protein(s) of interest. Therefore, different fixative methods are crucial in IIF staining of centromere-kinetochore proteins.
Here optimized methods of indirect immunofluorescent (IIF) staining and an assay to address localization of endogenous centromere-kinetochore proteins, including CENP-A and Flag-tagged exogenous CENP-A proteins, and quantitation of these proteins in human cells have been developed. These methods can be applied to the analysis of centromere-kinetochore proteins in other species.
Protocol
1. Cell Culture and Transfection
- Put a cover glass (22 mm x 22 mm) in a 6-well polystyrene plate. Optionally coat a cover glass with Poly-L-Lysine, 0.1% w/v, in water (see List of Materials/Equipment) to keep mitotic cells on the cover glass following the steps below:
Note: Optimal conditions must be determined for each cell line and application.- Aseptically coat culture surface with Poly-L-Lysine, 0.1% w/v, in water (0.4 ml/well of a 6-well polystyrene plate). Rock gently to ensure even coating of the culture surface.
- After 5 min, remove solution by aspiration and thoroughly rinse surface with sterile tissue culture grade water.
- Dry at least 2 hr before introducing cells and medium.
- Seed HeLa cells17 or HeLa Tet-Off cells17,22 on a cover glass (22 mm x 22 mm) put in a 6-well polystyrene plate. Check that cell density is 5.4 x 105 per well. Culture cells in high-glucose DMEM with 10% FBS and 1% penicillin-streptomycin.
Note: For optimal results, empirically determine the cell density to use in seeding. - Incubate the cells at 37 °C in an atmosphere of 5% CO2 for 18 hr.
Note: In the case of HeLa Tet-Off cells, transcription of the exogenous gene is active in the absence of the inducer (i.e., tetracycline/doxycycline) therefore culture cells without tetracycline/doxycycline and transfect transiently with the pTRM4 overexpression vector (Table 2), whose transcription is regulated by the TRE promoter (see below). - Eighteen hours after seeding, transfect cells as follows:
- Make solution A by mixing 1.5 µl siRNA oligo (20 µM annealed stock; Table 1) and/or 2.0 µg plasmid (Table 2) in 50 µl reduced serum medium (see List of Materials/Equipment), and incubate at RT for 5 min.
Note: In this analysis, CA-UTR siRNAs (a mixture of 5' and 3' UTR siRNA; Table 1) were co-transfected for use in Protocols 3 and 4 (see Discussion). - Make solution B by mixing 0.75 µl transfection reagent I (see List of Materials/Equipment) in 50 µl reduced serum medium, and incubate at RT for 5 min.
Note: An optional step is the addition of 1.0 µl transfection reagent II (see List of Materials/Equipment). - Mix solutions A and B together, and incubate at RT for 15 min.
- Wash the cultured cells once with PBS, and then add 500 µl reduced serum medium to each well of the 6-well polystyrene plate. Add the mixture of solutions A and B (i.e., RNA and /or DNA-lipid complex) directly to each of the individual well.
Note: Final concentration is 3.3 µg/ml (plasmid); 50 nM (siRNA). - Incubate the cells at 37 °C in an atmosphere of 5% CO2 for 4.5 hr. Change the medium to high-glucose DMEM with 10% FBS and 1% penicillin-streptomycin.
- Incubate the cells at 37 °C in an atmosphere of 5% CO2 for 48-72 hr after transfection.
Note: For optimal results, the incubation period for cell growth before fixation must be determined empirically, and protein depletion and/or expression must be confirmed by Western blot analysis (see Protocol 6). - If mitotic cell analysis is of interest, add paclitaxel (10 nM) to cultured cells 24 hr before fixation, or add TN16 (0.5 µM) to the cultured cells 2.5 hr before fixation.
- Make solution A by mixing 1.5 µl siRNA oligo (20 µM annealed stock; Table 1) and/or 2.0 µg plasmid (Table 2) in 50 µl reduced serum medium (see List of Materials/Equipment), and incubate at RT for 5 min.
2. Cell Fixation and Immunofluorescent Staining to Detect Endogenous Centromere-kinetochore Proteins (Paraformaldehyde Fixation)
- Preparation of fixative, buffers, and reagents.
- Freshly prepare 50 ml of 4% paraformaldehyde solution in PBS, pH 7.4.
- Add 40 ml of 1× PBS to a glass beaker on a stir plate in a ventilated hood. Heat while stirring to approximately 60 °C. Add 2 g of paraformaldehyde powder to the heated PBS.
- Raise slowly the pH by adding 1 ml of 1 N NaOH in total, because the powder will not immediately dissolve.
Note: The solution clears after the addition of NaOH. - Once paraformaldehyde has dissolved, cool and filter the solution.
- Adjust the pH with 1 N HCl to pH 7.4 and the volume of the solution with 1× PBS to 50 ml.
Note: Aliquots of the solution can be frozen or stored at 2-8 °C for as long as 1 week. The solution should be ice-cold or stored at 4 °C until it is to be used.
- Prepare 50 ml of buffer KB1, which consists of 10 mM Tris-HCl, pH 7.5; 150 mM NaCl; 0.5% BSA; and 0.5% Triton X-100.
Note: BSA should be freshly added.
Note: The buffer KB series are based on buffer KB described in previous reports.1,2,4 - Prepare 50 ml of buffer KB2, which consists of 10 mM Tris-HCl, pH 7.5; 150 mM NaCl; and 0.5% BSA.
Note: BSA should be freshly added. - Prepare 1 ml of buffer KB3 containing DAPI (50 ng/ml).
- Prepare 100 ml of mounting medium, which consists of 1 mg/ml p-phenylenediamine; 10% PBS, and 90% glycerol.
- Adjust the pH of 1× PBS to 9.8 with 1 N NaOH, dissolve p-phenylenediamine in the solution, and then add glycerol.
- Store 1 ml aliquots at -80 °C. Protect from light.
- Freshly prepare 50 ml of 4% paraformaldehyde solution in PBS, pH 7.4.
- Remove the culture medium by aspiration at 48-72 hr post transfection (see Protocol 1) for cell fixation. Rinse cells once with PBS. Apply PBS to the side of the culture wells to avoid disturbing the surface of the cells.
Note: The optimal time point for cell fixation must be determined empirically. - Fix the cells in 4% paraformaldehyde in PBS for 30 min at 4 °C. Rinse the cells twice with buffer KB2 to sufficiently remove residual 4% paraformaldehyde.
- Permeabilize the samples in buffer KB1 for 30 min at RT. Rinse the cells once with buffer KB2, and add buffer KB2 for 5 min at RT to additional block sites of nonspecific binding.
Note: Buffer KB1 also contributes significantly to blocking. - Remove a cover glass (22 mm x 22 mm) from a 6-well polystyrene plate using forceps. Using a hydrophobic barrier pen (see List of Materials/Equipment), draw a pale green square or circle to form a hydrophobic barrier around each cover glass sample. Do not touch or get too close to the cells with the hydrophobic barrier pen. Put the cover glass in new 6-well polystyrene plate.
- Dilute a primary antibody to centromere or kinetochore protein (dilution ratio of 1:100 to 1:200; see also List of Materials/Equipment) and either an anti-CENP-B antibody (dilution ratio of 1:400) or an anti-centromere antibody (ACA) (dilution ratio of 1:2,000) as a centromere location marker in buffer KB2.
Note: For optimal results, the final concentration of the primary antibody in this solution must be determined empirically. - Apply a sufficient volume (ca. 30 µl) of the diluted primary antibody to immerse the cell sample. Incubate the cell sample for 1 hr at 37 °C. Rinse the cells 3 times with buffer KB2.
- Using buffer KB2, dilute a fluorophore-conjugated secondary antibody (dilution ratio of 1:100 to 1:200) directed against each primary antibody.
Note: For optimal results, the final concentration of the secondary antibody in this solution must be determined empirically. - Apply a sufficient volume (a drop of ca. 30 µl on the cover glass) of the diluted secondary antibody to immerse the cell sample. Incubate the cell sample for 1 hr at 37 °C. Rinse the cells 5 times with buffer KB2 during a period of 30 min (five 6 min washes).
Note: For optimal results (i.e., for minimal loss of cells), the optimal washing condition must be determined empirically. - Apply a sufficient volume of buffer KB3 containing DAPI (50 ng/ml) to immerse the cell sample. Incubate the cell sample for 5 min at RT. Rinse the cells 1-2 times with buffer KB2.
- Mount the cover glass that contains the cell sample onto the micro slide.
- Place a drop of mounting medium in the center of the micro slide.
- Remove liquid from the cell sample and, using hands or forceps, position the sample in the center of the micro slide. Avoid air bubbles.
- Remove excess mounting medium with a paper towel.
3. Cell Fixation and Immunofluorescent Staining of C-terminal Flag-tagged CENP-A Proteins (Acetone Fixation)
- Preparation
- Prepare ice-cold 75% acetone.
- Freshly prepare 50 ml of PBS (pH 7.4) containing 0.5% skim milk and 0.5% BSA.
- Freshly prepare 50 ml of PBS (pH 7.4) containing 0.1% skim milk and 0.1% BSA.
- Prepare 1 ml of PBS (pH 7.4) containing DAPI (50-100 ng/ml).
- Prepare 100 ml of mounting medium as described in 2.1.5.
- Remove the culture medium by aspiration at 48-72 hr post transfection (see Protocol 1) for cell fixation. Rinse cells once with PBS. Apply PBS to the side of the culture well to avoid disturbing the surface of the cells.
Note: The optimal time point for cell fixation must be determined empirically. - Fix the cells in ice-cold 75% acetone, and incubate the cells for 10 min at -20 °C. Dry cells on the cover glass in a fume hood for 30-60 min at RT.
Note: For optimal results, the length of time needed for fixation and cell drying must be determined empirically. - Using the hydrophobic barrier pen (see List of Materials/Equipment), draw a pale green square or circle to form a hydrophobic barrier around each cover glass sample. Do not touch or get too close to the cells with the hydrophobic barrier pen.
- Block nonspecific binding sites on the cells by adding PBS containing 0.5% skim milk and 0.5% BSA for 5 min at RT.
- Using PBS containing 0.1% skim milk and 0.1% BSA, dilute an anti-Flag antibody (1:1,000 dilution ratio) and either an anti-CENP-B antibody (dilution ratio of 1:200) or ACA (1:2,000 dilution ratio) as a centromere location marker.
Note: For optimal results, the final concentration of the primary antibody in this solution must be determined empirically. - Apply a sufficient volume (ca. 30 µl) of the diluted primary antibody to immerse the cell sample. Incubate the cell sample for 1 hr at 37 °C. Rinse the cells 5 times with the blocking buffer during a period of 30 min.
Note: Cells can be rinsed with PBS. For optimal results (i.e., to avoid loss of cells), the optimal washing conditions must be determined empirically. - Dilute a fluorophore-conjugated secondary antibody (dilution ratio of 1:100 to 1:200) directed against the each primary antibody in PBS containing 0.1% skim milk and 0.1% BSA.
Note: For optimal results, the final concentration of the secondary antibody in this solution must be determined empirically. - Apply a sufficient volume (ca. 30 µl) of the diluted secondary antibody to immerse the cell sample. Incubate the cell sample for 1 hr at 37 °C. Rinse the cells 2 times with PBS containing 0.1% skim milk and 0.1% BSA.
Note: PBS alone can also be used to wash the cells. - Apply a sufficient volume of PBS containing DAPI (50-100 ng/ml) to immerse the cell sample. Incubate the cell sample for 5 min at RT. Rinse the cells 1-2 times with PBS.
- Mount the cover glass containing the cell sample onto the micro slide as described in 2.11.
4. Cell Fixation and Immunofluorescent Staining of N-terminal Flag-tagged CENP-A Proteins (Methanol Fixation)
- Preparation
- Prepare ice-cold methanol.
- Prepare TBS (pH 7.4) containing 4% goat serum.
- Prepare 1 ml of TBS (pH 7.4) containing DAPI (50-100 ng/ml).
- Prepare 100 ml of mounting medium as described in 2.1.5.
- Remove the culture medium by aspiration at 48-72 hr post transfection (see Protocol 1) for cell fixation. Rinse cells once with TBS. Apply TBS to the side of the culture wells to avoid disturbing the surface of cells.
Note: The optimal time point for cell fixation must be determined empirically. - Fix the cells in ice-cold methanol, and incubate the cells for 6 min at -20 °C. Rinse the cells twice with TBS to sufficiently remove residual methanol.
- Using a hydrophobic barrier pen (see List of Materials/Equipment), draw a pale green square or circle to create a hydrophobic barrier around each cover glass sample. Do not touch or get too close to the cells with the hydrophobic barrier pen.
- Block nonspecific binding sites on the cells by adding TBS containing 4% goat serum. Incubate for 10 min at RT.
- Dilute an anti-Flag antibody (1:1,000 dilution) and either an anti-CENP-B antibody (dilution ratio of 1:200) or ACA (1:2,000 dilution ratio) as a centromere location marker in TBS containing 4% goat serum.
Note: For optimal results, the final concentration of the primary antibody in this solution must be determined empirically. - Apply a sufficient volume (ca. 30 µl) of the diluted primary antibody to immerse the cell sample. Incubate the cell sample for 1 hr at 37 °C. Rinse the cells 5 times with the blocking buffer during a period of 30 min. For optimal results (i.e., the minimal loss of cells), the optimal washing condition must be determined empirically.
- Dilute a fluorophore-conjugated secondary antibody (dilution ratio of 1:100 to 1:200) directed against each primary antibody in TBS containing 4% goat serum.
Note: For optimal results, the final concentration of the secondary antibody in this solution must be determined empirically. - Apply a sufficient volume (ca. 30 µl) of the diluted secondary antibody to immerse the cell sample. Incubate the cell sample for 1 hr at 37 °C. Rinse the cells 3 times with the blocking buffer.
- Apply a sufficient volume of TBS containing DAPI (50-100 ng/ml) to immerse the cell sample. Incubate the cell sample for 5 min at RT. Rinse the cells 1-2 times with TBS.
- Mount the cover glass that contains the cell sample onto the micro slide as described in 2.11.
5. Immunofluorescence Image Observation, Acquisition, Quantitation, and Analysis
- Observe the cell sample through a motorized fluorescence microscope equipped with an 63X and 100X oil immersion lens, an external compact light source, and a digital CCD camera.
- Perform image acquisition and processing, including deconvolution, by using Software A, or Softwares B1 and B2 (see List of Materials/Equipment). Please see Supplemental Code Files (5.2.1) for all commands used in Software A. For all commands used in Softwares B1 and B2, please see (5.2.2) in Supplemental Code Files.
- Use a previously described method23-25 to quantify signals of centromere-kinetochore proteins (e.g., remaining signals of CENP-A at the centromere) with the following minor modifications:
- Select area of the centromere-kinetochore proteins and that of the background as follows:
- Mitotic cell: (centromere-kinetochore region) Select area overlapping with chromosomes stained with DAPI; (background region) and area outside chromosomes but inside the identical single cell (i.e., cytosolic region).
- Interphase cell: (centromere-kinetochore region) Select area overlapping with chromatin stained with DAPI; (background region) and area outside chromatin but inside the identical single cell (i.e., cytosolic region).
Note: See also Supplemental Code Files (5.2.1.4.3) or (5.2.2.6.4) for area selection with Software A or Software B2, respectively.
- Quantify the percentage of remaining signals at the centromeres by using Software A or B. For this task, use the following formula:
Remaining signals of centromere-kinetochore protein at the centromere-kinetochore

where s is the signal brightness of the selected area, which is confirmed by ACA or CENP-B staining; b is the background signal brightness; rsample is the reference ACA or CENP-B signals for siRNA(s)-treated cells; and rctrl is the reference ACA or CENP-B signals for Luc siRNA-transfected cells.
Note: In this analysis, CENP-B signals were used as reference signals for CUL4A and RBX1 as described in previous reports.17- Use an Excel file for this calculation after copying and pasting the raw data from the signal quantitation software described above. See also Supplemental Code Files: (5.2.1.4) and (5.2.2.6) for details. One example of calculation is shown in Table 3.
- Analyze at least 20 cells to eliminate variation in staining and image acquisition for each measurement level. Optionally use "averaged value" of remaining centromere-kinetochore signals for each analyzed cell to compare these values among different centromere-kinetochore proteins.
- Select area of the centromere-kinetochore proteins and that of the background as follows:
6. Western Blot Analysis of Total Protein
- Resuspend the cells in denaturing buffer A (20 mM Tris-HCl, pH 7.4; 50 mM NaCl; 0.5% Nonidet P-40; 0.5% deoxycholate; 0.5% SDS; 1 mM EDTA; and complete EDTA-free protease inhibitor cocktail),26 subject the suspension to a sonication and freeze-thaw process, and measure protein concentrations as follows:
- For the sonication process, add 50 µl of buffer A to the cells collected from two wells of a 6-well polystyrene plate at 48-72 hr post transfection (see Protocol 1). Operate a sonicator equipped with disruptor horn and microtip (see List of Materials/Equipment) for total 15 sec intermittent-pulse duration (duty cycle 50%) per one sample.
- For freeze-thaw process, freeze cells with liquid nitrogen and thaw cells at RT.
- Measure protein concentrations using a commercial protein assay reagent I or II (see List of Materials/Equipment).
Note: Lysates are diluted with ratio of 1:10 in the measurement of protein concentrations with either reagent I or II. At this dilution, the SDS present in buffer A shows little or no interference in this measurement.
- Mix the lysate containing 20-30 µg of total protein with 2× or 4× SDS-PAGE loading buffer.27 Boil the samples for 5 min and then load them on a 12.0%-15.0% denaturing SDS-polyacrylamide gel for electrophoresis.
- Transfer the proteins separated by SDS-PAGE onto a PVDF membrane by using a Western blotting method described previously.17,24,27-31
- Block the nonspecific binding sites on the membrane with 5% non-fat milk in 1× PBS, and then incubate the membrane with solutions of diluted primary antibodies for 1 hr at RT. See List of Materials/Equipment for detailed information (e.g., dilution ratio) of each primary antibody.
- After washing the membrane 3-4 times (each 3- to 5-min incubation with shaking) with PBS-T buffer (1× PBS and 0.1% Tween-20), incubate the membrane in a mixture of near-infrared (IR) fluorescent dye-conjugated secondary antibodies (dilution ratio of 1:20,000), DyLight-conjugated secondary antibodies (dilution ratio of 1:20,000), and/or horseradish peroxidase (HRP)-conjugated secondary antibodies (diluted in PBS-T; dilution ratio of 1:10,000) for 1 hr at RT. See List of Materials/Equipment for detailed information (e.g., dilution ratio) of each secondary antibody.
- Wash the membrane 3 times, and then scan the membrane to analyze the proteins with the infrared imaging system and/or the chemiluminescence imager for immunoblot detection (see List of Materials/Equipment).
- For using the infrared imaging system, see (6.4.1) in Supplemental Code Files.
- For using the chemiluminescence imager, see (6.4.2) in Supplemental Code Files. Use an ultra-sensitive enhanced chemiluminescent (ECL) substrate (see List of Materials/Equipment) for this system.
Representative Results
Immunofluorescence analysis of endogenous CENP-A supports the hypothesis that CUL4A-E3 ligase is required for localization of CENP-A to centromeres
Our recent studies showed that CUL4A-RBX1-COPS8 E3 ligase activity is required for ubiquitylation of lysine 124 (K124) on CENP-A and localization of CENP-A to centromeres.17 Initially, the interphase-centromere complex (ICEN) was isolated by anti-CENP-A:native chromatin immunoprecipitation,32-34 and it has been hypothesized that some of the ICEN proteins may play a role in localizing CENP-A to centromeres. Therefore, siRNA knockdown experiments were performed to screen for proteins whose absence of expression induced the delocalization of CENP-A at centromeres17 (data not shown): asynchronously growing HeLa cells were transfected with Cullin 4A (CUL4A) siRNA or RBX1 siRNA for the times required for optimal protein depletion (48 hr for CUL4A and 72 hr for RBX1). Cells transfected with CUL4A siRNA showed a significant delocalization of CENP-A at centromeres (Figure 1A-C). As seen in Figure 2G, the protein levels of CENP-A in total cell lysates of CUL4A siRNA-transfected cells were similar to CENP-A levels in lysates from luciferase (Luc) siRNA-transfected cells under the same culture conditions. The possibility of off-target effects of CUL4A siRNA was excluded, because ectopic expression of CUL4A-Flag rescued the reduction of CENP-A at centromeres when CUL4A siRNA targeted the 3' UTR (Figure 2A-C). The protein levels of CENP-A in total cell lysates were confirmed to be similar to CENP-A levels in lysates from Luc siRNA-transfected cells under the same culture conditions regardless of the cell cycle stage (Figures 1B, 2G); thus, the possibility that CUL4A depletion caused CENP-A protein degradation was eliminated.
Cullin-RING-E3 ubiquitin ligases (CRLs) are the most prominent class of ubiquitin ligases35 and contain 3 major elements: a cullin scaffold, a RING finger protein (RBX1 or RBX2), and a ubiquitin-charged E2 enzyme that is recruited by RBX1 or RBX2,36 A RING finger protein also recruits a substrate adaptor that places substrates in proximity to the E2 enzyme to facilitate ubiquitin transfer.36 RBX1 siRNAs induced a significant reduction of CENP-A at centromeres (Figure 1D-F) under the conditions in which the protein levels of CENP-A in total cell lysates of RBX1 siRNA-transfected cells were similar to CENP-A levels in lysates from Luc siRNA-transfected cells (Figure 2G). Degradation of CENP-A protein either by RBX1 depletion or by the combination of CUL4A and RBX1 depletion under the same culture conditions was not observed (Figure 2G). The possibility of off-target effects of RBX1 siRNA was excluded, because ectopic expression of Flag-RBX1 rescued the reduction of CENP-A at centromeres when RBX1 siRNA targeted the 3' UTR (Figure 2D-F). These findings suggested that CUL4A-RBX1-E3 ligase is specifically required for the localization of CENP-A to centromeres.
Immunofluorescence analysis of exogenous CENP-A–Flag proteins indicates that CENP-A K124 ubiquitylation is essential for localization of CENP-A to centromeres
Previously, our immunoprecipitation-mass spectrometry analysis showed that lysine 124 (K124) of CENP-A-Flag is ubiquitylated in HeLa cells.17 In the crystal structure of the CENP-A nucleosome, K124 resides in the α3 helix, although the site is not within the CATD region.17 In addition, K124 is conserved among mammals, birds, lizards, plants, and a group of fungi (e.g., budding yeast).17 CENP-A lysine mutants (Figure 3A) were constructed and tested their ability to localize to the centromere. Substantial abrogation of the centromeric localization of exogenous CENP-A K124R mutant with diffuse signals in both mitotic and interphase HeLa cells was observed (Figure 3B, C). Centromere localization was not significantly affected neither by K9A mutations (K9 corresponds to histone H3 K9 methylation) nor K77R mutations (K77 is a unique lysine site in CATD) (Figure 3B, C).
Based on these results, two hypotheses are proposed regarding the in vivo function of CENP-A ubiquitylation at K124. First, the role of K124 ubiquitylation is for ubiquitin-mediated proteolysis to eliminate overexpressed and/or mislocalized CENP-A to euchromatin. Second, the role of CENP-A ubiquitylation on K124 is for loading CENP-A onto centromeres. To test the first possibility, the stabilities of CENP-A-Flag wild type and the K124R mutant as well as endogenous CENP-A after cycloheximide (CHX) treatment of CUL4A- or RBX1-depleted cells were addressed. These data suggested that ubiquitylation of K124 is probably not involved in ubiquitin-mediated proteolysis to eliminate overexpressed and/or mislocalized CENP-A (data not shown).17. Furthermore, it was confirmed that the K124R mutation abrogates putative monoubiquitylation and diubiquitylation bands, both in in vivo and in vitro ubiquitylation assays (data not shown).17 Collectively, these data suggest that the CUL4A-RBX1 complex contributes to "signaling" ubiquitylation, which is required for CENP-A localization at centromeres.
Immunofluorescence analysis of exogenous Flag–CENP-A proteins indicates that monoubiquitin fusion is sufficient to load CENP-A K124R at centromeres
Based on the results above, it was hypothesized that covalently linked monoubiquitin serves as a signal for CENP-A loading onto centromeres. To test this hypothesis, an N-terminal Flag-tagged and C-terminal ubiquitin-fused wild-type CENP-A and a K124R mutant CENP-A was constructed (Figure 4A). The monoubiquitin mutant Ub (K48R), which lacks a major site for polyubiquitylation37-39 was also used to prevent ubiquitin-fused CENP-A protein from potential polyubiquitylation. By capturing anti-Flag immunofluorescent signals, the centromeric localization of proteins encoded by these constructs was tested. Whereas Flag-CENP-A (K124) substantially abrogated centromere localization of CENP-A (Figure 4B and 4C, column [6]), both Flag-CENP-A (WT) and Flag-CENP-A (WT)-Ub (WT) maintained their centromere localization (Figure 4B and 4C, columns [1] and [2]). The Flag-CENP-A (K124R)-Ub (K48R) protein presumably mimicked monoubiquitylated CENP-A, as this protein substantially restored localization to centromeres (Figure 4C, compare columns [5] and [6]) more efficiently than did CENP-A (K124R)-Ub (WT) (Figure 4C, compare columns [4]-[6]). These data demonstrated that monoubiquitylation is sufficient for recruitment of CENP-A to centromeres.

Figure 1. Immunofluorescence analysis of endogenous CENP-A supports the hypothesis that CUL4A-E3 ligase is required for localization of CENP-A to centromeres (Figures were adapted from Niikura et al.17). (A) CUL4A siRNA induced delocalization of CENP-A at centromeres. HeLa cells were transfected with CUL4A or luciferase (Luc) siRNA and incubated for 48 hr (Table 1). Scale bar represents 10 μm. (B) Western blot analysis of HeLa whole cell lysates using the same culture condition as in (A). Cells were harvested 48 hr after transfection with CUL4A siRNA or Luc siRNA (Table 1). GAPDH served as a loading control. (C) Quantified endogenous CENP-A signals at centromeres shown in (A). Normalization of signals was performed by using Luc siRNA-transfected cells, and the mean percentages (±SD) are shown. ****P < 0.0001 vs Luc siRNA-transfected cells (Student's t-test). (D) RBX1 siRNA induced delocalization of CENP-A from centromeres. HeLa cells were transfected with RBX1 or Luc siRNA(s) and incubated for 72 hr (Table 1). Scale bar represents 10 μm. (E) Western blot analysis of HeLa whole cell lysates using the same culture condition as in (D). Cells were harvested 72 hr after transfection with RBX1 or Luc siRNA(s) (Table 1). GAPDH served as a loading control. (F) Quantified endogenous CENP-A signals at centromeres shown in (D). Normalization of signals was performed by using Luc siRNA-transfected cells, and the mean percentages (±SD) are shown. ****P < 0.0001 vs Luc siRNA-transfected cells (Student's t-test). Please click here to view a larger version of this figure.

Figure 2. Supplemental information related to Figure 1 centromeres (Figures were adapted from Niikura et al.17). (A) Exogenous CUL4A-Flag rescued the delocalization of CENP-A when CUL4A siRNA targeted the 3' UTR. HeLa Tet-Off cells were fixed at 48 hr after cotransfection with siRNA (CUL4A #2: 3' UTR target or Luc; Table 1) plus the plasmid construct (pTRM4-CUL4A-Flag or vector; Table 2). 4-color immunostaining: staining for DAPI (blue); Flag; endogenous CENP-B (red); and endogenous CENP-A (green), was performed and Flag-positive cells were sorted, but Flag images are omitted for simplicity. Note: CUL4A #2 is the identical target as that shown in Figure 1A-C. Scale bar represents 10 μm. (B) Western blot analysis of HeLa Tet-Off whole cell lysates following the same culture condition as shown in (A). GAPDH served as a loading control. (C) CENP-A signals at centromeres shown in (A) were quantified by microscopy after the sorting of cells to isolate those bound by an anti-Flag antibody. Normalization of signals was performed with cells transfected with Luc siRNA plus vector, and the mean percentages (±SD) are shown. ****P < 0.0001 vs cells transfected with Luc siRNA plus vector (left column, Student's t-test). (D) Exogenous Flag-RBX1 rescued the delocalization of CENP-A at the centromere when RBX1 siRNA targeted the 3' UTR. HeLa cells were fixed at 72 hr after cotransfection with siRNA (RBX1 #2: 3' UTR target or Luc; Table 1) plus the plasmid construct (pcDNA3-Flag-RBX1 or vector; Table 2). 4-color immunostaining: staining for DAPI (blue); Flag; endogenous CENP-B (red); and endogenous CENP-A (green), was performed and Flag-positive cells were sorted, but Flag images are omitted for simplicity. Scale bar represents 10 μm. (E) Western blot analysis of HeLa whole cell lysates following the same culture condition as in (D). GAPDH served as a loading control. (F) CENP-A signals at centromeres shown in (D) were quantified by microscopy after the sorting of cells to isolate those bound by an anti-Flag antibody. Normalization of signals was performed with cells transfected with Luc siRNA plus vector, and the mean percentages (±SD) are shown. ****P < 0.0001 vs cells transfected with Luc siRNA plus vector (left column, Student's t-test). (G) Depletion of CUL4A and RBX1 did not affect levels of endogenous CENP-A protein. Levels of endogenous CENP-A protein were measured in HeLa whole cell lysates harvested 72 hr after transfection with CUL4A, RBX1, CUL4A plus RBX1, or Luc siRNA. Forty-eight hours after transfection, cells were either untreated (Asyn) or treated with paclitaxel (+Tax) to induce arrest in mitosis, and they were cultured for 24 hr. GAPDH served as a loading control. Please click here to view a larger version of this figure.

Figure 3. Immunofluorescence analysis of exogenous CENP-A-Flag proteins indicated that CENP-A K124 ubiquitylation is required for CENP-A localization at centromeres (Figures were adapted from Niikura et al.17). (A) Overexpression of CENP-A-Flag constructs (WT and KR mutants) was confirmed by Western blot analysis of HeLa Tet-Off whole cell lysates. Cells were harvested 48 hr after transfection with pTRM4-CENP-A-Flag WT, KR mutants, or pTRM4 vector (Table 2). Overexpression of CENP-A-Flag was detected by an anti-Flag antibody. GAPDH served as a loading control. Putative CENP-A-Flag dimer (##) and CENP-A-Flag monomer (#) are shown. Note: SDS-resistant CENP-A dimers have been reported previously.40,41 (B) The CENP-A K124R mutant showed diffused delocalization from centromeres. Signals from DAPI (blue), Flag (green), and endogenous CENP-B (red; a centromere location control) are shown. Note: Unlike endogenous CENP-A in cells transfected with CUL4A or RBX1 siRNA, diffuse signals appeared in cells that overexpressed exogenous CENP-A K124R-Flag as a phenotype of the inability to be targeted to centromeres, presumably because its expression level is approximately 10- to 25-fold higher than that of endogenous CENP-A (data not shown).17 Scale bar represents 10 μm. (C) Histograms of the localization patterns shown in (B). More than 50 pro/prometaphase and metaphase cells and more than 200 interphase cells per experiment were counted (n ≥ 3 experiments). The mean percentages (±SD) are shown. "Others (Non-centromere)" indicates mostly damaged cells, dead cells, or cells with nucleolar localization (only in interphase), presumably because of transfection or other treatments. ***P < 0.001 vs CENP-A WT-Flag (Student's t-test). Please click here to view a larger version of this figure.
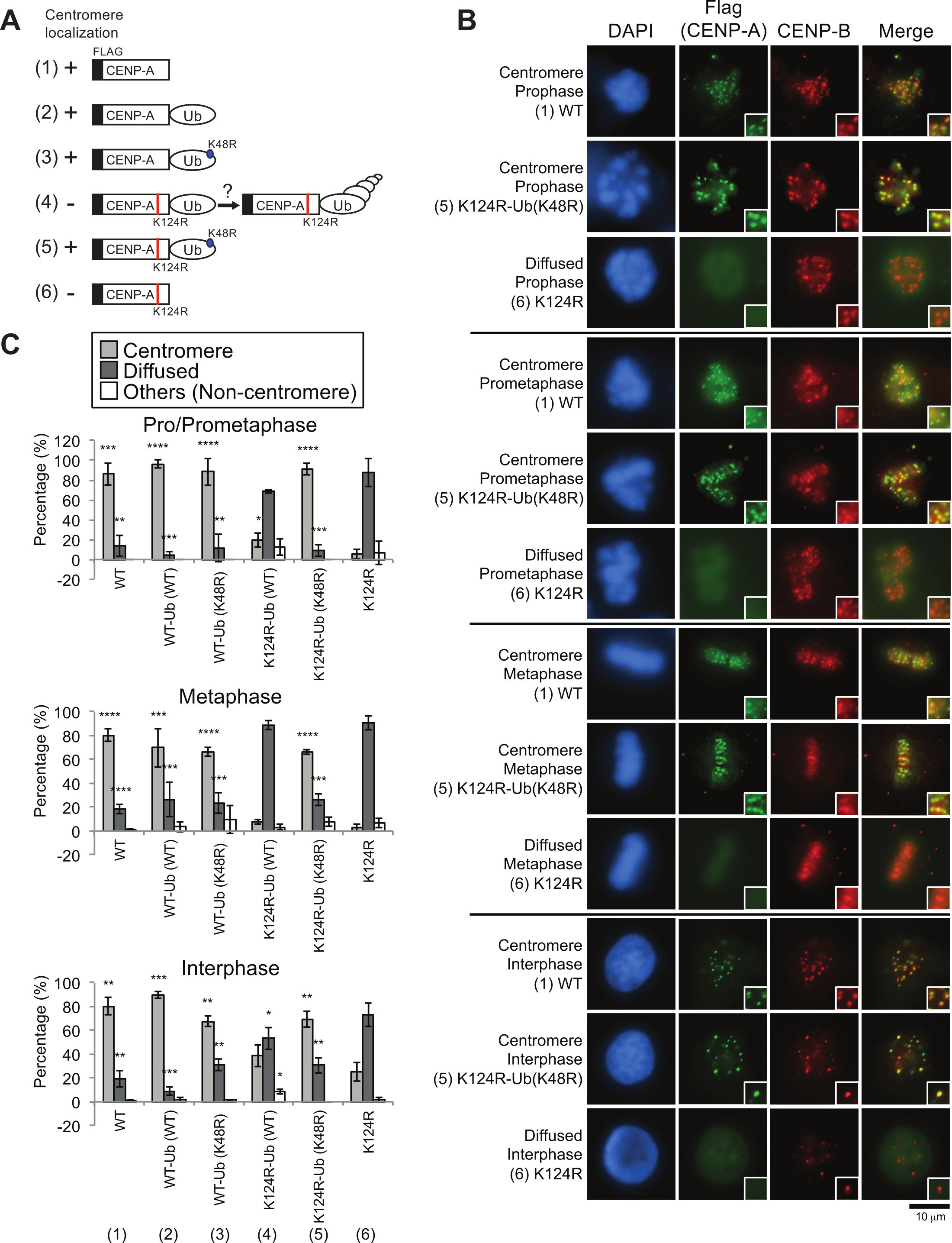
Figure 4. Immunofluorescence analysis of exogenous Flag-CENP-A proteins indicated that CENP-A monoubiquitin fusion rescued delocalization of the CENP-A K124R mutant from centromeres (Figures were adapted from Niikura et al.17). (A) Schematic cartoons of each construct used in this study (see also Table 2). The K124R mutation site on CENP-A (red) and the K48R mutation site on monoubiquitin (blue) are shown. (1) WT: Flag-CENP-A WT, (2) WT-Ub (WT): Flag-CENP-A WT-Ub (WT), (3) WT-Ub (K48R): Flag-CENP-A WT-Ub (K48R), (4) K124R-Ub (WT): Flag-CENP-A K124R-Ub (WT), (5) K124R-Ub (K48R): Flag-CENP-A K124R-Ub (K48R), (6) K124R: Flag-CENP-A K124R. (B) Exogenous CENP-A monoubiquitin fusion rescued delocalization of the CENP-A K124R mutant from centromeres. DAPI (blue), Flag (green), and endogenous CENP-B (red; a centromere location control) are shown. Scale bar represents 10 µm. (C) Histograms of the localization patterns shown in (C). More than 50 pro/prometaphase and metaphase cells and more than 200 interphase cells per experiment were counted (n ≥ 3 experiments). The mean percentages (±SD) are shown. ****P < 0.0001 and ***P < 0.001 vs non-fused Flag-CENP-A K124R (column [6]) (Student's t-test). Please click here to view a larger version of this figure.
| siRNA target | Indication in present study | Target type | siRNA database number | Forward sequence(s) | Source / Reference | |
| Luciferase (GL3) | - | 1 target | RKK9 | CUUACGCUGAGUACUUCGAdTdT | Elbashir et al., (2001) | |
| CENP-A | CA-UTR | siRNA pool (2 targets mixture) | RKK375 / 5' UTR t1 | CGAGCGGCGCGGACUUCUGCCdTdT | Niikura et al., (2015) | |
| RKK383 / 3' UTR t1 | UCCUGCACCCAGUGUUUCUGUdGdT | Niikura et al., (2015) | ||||
| CUL4A | #1 | 1 target | CRKK178 / RKK309 / t1 | AGCGAUCGUAAUCAAUCCUGA | Niikura et al., (2015) | |
| #2 (3'UTR) | 1 target | CRKK181 / RKK331 / t4 | AUGCGGGUUUGAAAUAUGACA | Niikura et al., (2015) | ||
| RBX1/ROC1 | #1 | siRNA pool (4 targets mixture) | CRKK198 / RKK411 / t1 | GAAGCGCUUUGAAGUGAAA | Niikura et al., (2015) | |
| CRKK198 / RKK413 / t2 | GCAUAGAAUGUCAAGCUAA | Niikura et al., (2015) | ||||
| CRKK198 / RKK415 / t3 | GCAAGAAGCGCUUUGAAGU | Niikura et al., (2015) | ||||
| CRKK198 / RKK417 / t4 | CAACAGAGAGUGGGAAUUC | Niikura et al., (2015) | ||||
| #2 (3'UTR) | 1 target | CRKK206 / RKK425 / t8 | UUCCCUGCUGUUACCUAAUUA | Niikura et al., (2015) | ||
Table 1. siRNA sequences used in this study.
| B number | Relevant characteristic(s) | Source/Reference |
| B288 | pTRM4 | Niikura et al., (2006) |
| B2067 | pTRM4-human CENP-A-Flag | Niikura et al., (2015) |
| B2281 | pTRM4-human CENP-A K9A-Flag | Niikura et al., (2015) |
| B2387 | pTRM4-human CENP-A K77R-Flag | Niikura et al., (2015) |
| B2388 | pTRM4-human CENP-A K124R-Flag | Niikura et al., (2015) |
| B2512 | pTRM4-Flag-human CENP-A | Niikura et al., (2015) |
| B2579 | pTRM4-Flag-human CENP-A K124R | Niikura et al., (2015) |
| B2513 | pTRM4-Flag-human CENP-A-Ub (WT) | Niikura et al., (2015) |
| B2559 | pTRM4-Flag-human CENP-A K124R-Ub (WT) | Niikura et al., (2015) |
| B2515 | pTRM4-Flag-human CENP-A-Ub (K48R) | Niikura et al., (2015) |
| B2560 | pTRM4-Flag-human CENP-A K124R-Ub (K48R) | Niikura et al., (2015) |
| B2759 | pTRM4-human CUL4A-Flag | Niikura et al., (2015) |
| B2624 | pcDNA3-Flag-human RBX1 | Niikura et al., (2015) |
| B1491 | pcDNA3-Flag | Niikura et al., (2015) |
Table 2. Plasmid vectors used in this study.
| R (anti-CENP-B) sample | b (anti-CENP-B) | R sample (subtracted) | Ratio (S sample : R sample) | Corrected Ratio (S sample : R sample) |
| 520 | 514 | 6 | -4.167 | 0.000 |
| 535 | 445 | 90 | -1.033 | 0.000 |
| 515 | 431 | 84 | 0.274 | 0.274 |
| 562 | 483 | 79 | 0.506 | 0.506 |
| 902 | 562 | 340 | 0.509 | 0.509 |
| 1203 | 703 | 500 | -0.560 | 0.000 |
| 1014 | 589 | 425 | -0.819 | 0.000 |
| 768 | 510 | 258 | -1.345 | 0.000 |
| 555 | 458 | 97 | -1.845 | 0.000 |
| 781 | 576 | 205 | -1.146 | 0.000 |
| 556 | 436 | 120 | 0.758 | 0.758 |
| 534 | 493 | 41 | -1.634 | 0.000 |
| 702 | 543 | 159 | 0.667 | 0.667 |
| 482 | 429 | 53 | -1.000 | 0.000 |
| 654 | 531 | 123 | 0.740 | 0.740 |
| 1190 | 607 | 583 | 0.384 | 0.384 |
| 552 | 454 | 98 | 0.969 | 0.969 |
| 489 | 413 | 76 | 0.539 | 0.539 |
| 511 | 452 | 59 | -3.186 | 0.000 |
| 485 | 475 | 10 | -2.700 | 0.000 |
| 481 | 441 | 40 | -1.475 | 0.000 |
| Sum | 5.347 | |||
| Average | 0.255 | |||
| R (anti-CENP-B) ctrl | b (anti-CENP-B) | R ctrl (subtracted) | Ratio (S ctrl : R ctrl) | Corrected Ratio (S ctrl : R ctrl) |
| 543 | 483 | 60 | 3.017 | 3.017 |
| 526 | 466 | 60 | 2.667 | 2.667 |
| 532 | 460 | 72 | 1.792 | 1.792 |
| 507 | 457 | 50 | 3.520 | 3.520 |
| 491 | 458 | 33 | 4.273 | 4.273 |
| 545 | 464 | 81 | 2.160 | 2.160 |
| 518 | 484 | 34 | 3.559 | 3.559 |
| 461 | 404 | 57 | 2.404 | 2.404 |
| 487 | 447 | 40 | 3.550 | 3.550 |
| 534 | 477 | 57 | 3.965 | 3.965 |
| 528 | 456 | 72 | 3.097 | 3.097 |
| 707 | 601 | 106 | 1.868 | 1.868 |
| 615 | 528 | 87 | 2.828 | 2.828 |
| 660 | 481 | 179 | 1.620 | 1.620 |
| 565 | 476 | 89 | 1.607 | 1.607 |
| 527 | 455 | 72 | 3.583 | 3.583 |
| 560 | 474 | 86 | 2.930 | 2.930 |
| 510 | 451 | 59 | 4.017 | 4.017 |
| 729 | 586 | 143 | 2.678 | 2.678 |
| 634 | 576 | 58 | 3.655 | 3.655 |
| 507 | 476 | 31 | 3.935 | 3.935 |
| Sum | 62.725 | |||
| Average | 2.987 | |||
| Remaining CENP-A signals at centromere (%) | 8.5 |
Table 3. One example of calculation of remaining CENP-A signals at the centromere (%). An example of Pro/Prometaphase in Figure 2F is shown: sample is RBX1 #2 (3'UTR) + Vec (center column in Figure 2F) and control is Luc + Vec (left column in Figure 2F). As a result, remaining CENP-A signals at centromere (%) = (0.255 / 2.987) x 100 = 8.5% is obtained (Or [5.347 / 62.725] x 100 = 8.5% with the same total cell number [i.e., 21 cells] analyzed between two samples). Note that if b value is more than S value (i.e., if subtracted value is negative), corrected ratio value is set to 0.00.
Discussion
In recent years many studies have developed different quantitative microscopy assays for fixed cells.42 Progress in centromere-kinetochore biology often requires an understanding of the centromere-specific or kinetochore-specific function of proteins of which subcellular spatial-temporal regulation reflects the changing functions of these proteins during cell cycle. Therefore, here we developed methods of IIF staining and a quantitative IIF assay to specifically analyze the relative levels of endogenous and exogenous CENP-A proteins, which can be applicable in differently treated samples. Currently we achieved successful IIF staining of endogenous CENP-I, CENP-H, KNL1, Hec1, and Ska1 using Protocol 2 in this study (see Niikura et al.24 and data not shown; List of Materials/Equipment for information of antibodies). In future, one can apply the same method to quantify the level of different centromere-kinetochore proteins (both endogenous and Flag-tagged exogenous proteins) choosing the specific antibodies, however, our IIF methods do not cover the detection of these proteins in living cells or in a specific single cell during whole cell cycle. Because these procedures require cells to be fixed and processed on separate coverslips, we introduced reference signals (e.g., ACA or CENP-B) for the purpose of normalization in order to fairly compare signals among different samples and increase the accuracy of the assay. Because ACA involve endogenous and/or exogenous CENP-A signals, CENP-B as a reference signal for the purpose of normalization would be more appropriate for comparison than would ACA in terms of the accuracy of the quantitative assay of CENP-A signals. The amino-terminal region of CENP-B binds to the 17-bp motif of the CENP-B box sequence in alphoid DNA, and the carboxy-terminal region of CENP-B forms homodimers.43,44 CENP-B does not colocalize completely with CENP-A and/or other centromere-kinetochore proteins, however, is closely associated with them.45 While immunofluorescence with anti-CENP-C antibodies typically gives two discrete spots, staining with anti-CENP-B often appears as a single bright bar connecting both sister centromeres. 46 However, in our macro-scale microscopic observation, some CENP-B signals apparently overlap with CENP-A signals, especially at earlier mitotic stages (prophase-prometaphase than late metaphase), also in part depending on the observational angle and the type of fixative used (data not shown). In contrast, in a quantitative IIF assay of CENP-A signals, protein localization of reference signals should not be affected by depletion and/or dysfunction of CENP-A for fair analysis, although localization of most of the centromere-kinetochore proteins is dependent on CENP-A localization at centromeres.47,48 CENP-A-independent kinetochore assembly is established by replacing the DNA-binding regions of CENP-C and CENP-T with alternative chromosome-targeting domains that recruit these proteins to ectopic loci.49,50 In addition, recent work indicated that the inner kinetochore components CENP-C and CENP-T act in parallel to recruit the KMN network to kinetochores.51-54 Furthermore, Nishino et al. suggested that the CENP-T-W-S-X complex forms a unique nucleosome-like structure to generate contacts with DNA, extending the 'histone code' beyond canonical nucleosome proteins.55 Because the centromere localization of CENP-C is dependent on the centromere localization of CENP-A,47,48 CENP-T-W-S-X or especially CENP-T could be another protein candidate(s) to provide reference signals in a quantitative IIF assay of CENP-A. However, the candidate for reference signaling should be carefully tested before the assay to determine whether localization at centromeres is affected by depletion and/or dysfunction of CENP-A. Analogous consideration should be taken for quantitative IIF assays of other centromere-kinetochore proteins: protein localization of reference signals should not be affected by depletion and/or dysfunction of the target protein for fair analysis. In the previous study, we used ACA as a reference signal for quantitative IIF assay of other central-outer kinetochore proteins.24 ACA could be a more appropriate option than single CENP-B as a position control as well as a reference signal, if ACA signals are not affected by depletion and/or dysfunction of the target protein but colocalization of the target protein with CENP-B is poor as mentioned above.
Bodor et al. reported that centromeric CENP-A levels are regulated by mass action, i.e., the amount of centromeric CENP-A varies in direct proportion to the cellular content.56 They also showed that transient induction of CENP-A expression leads to a prompt increase in the centromeric CENP-A level. Conversely, we observed a large cell population with centromere-localized, Flag-tagged CENP-A WT after cotransfection of the pTRM4 expression vector with CA-UTR siRNAs (a mixture of 5' and 3' UTR siRNA; Table 1) (data not shown); this population was larger than that seen after transfection with only the pTRM4 expression vector. The level of exogenous Flag-tagged CENP-A protein whose expression was driven by pTRM4, was approximately 1.0 to 1.4 orders of magnitude (10- to 25-fold) higher than that of endogenous CENP-A (data not shown). It is speculated that expression of exogenous CENP-A WT above a particular threshold could adversely affect its proper centromeric localization.
In the present protocols, fixation is a critical step to maintain cellular structure and environment as proximal situation as possible to the native state. Once cells are fixed, one should proceed with the staining protocol to avoid loss of the protein of interest or the rest of the cell. However, fixation destroys antigenic sites occasionally, and different antibody-antigen combinations work poorly with one fixative, but very well with another.21 Because of this variation, the choice of fixative depends largely on the protein(s) of interest. The time length of fixation can significantly affect the detection of the protein(s) by immunostaining, and shorter fixation is generally better to retain antigenicity.57 Therefore, when determining staining procedures for a new target of other centromere-kinetochore proteins, especially of uncertain localization, or when working with a new antibody, one should test several methods of fixation and buffers to find the optimal condition. In the present protocols, two classes of popular fixatives were selected: aldehyde fixatives (Protocol 2) and organic solvents (Protocols 3 and 4). The purity of the fixatives is also extremely important. In Protocol 4, 4% goat serum is used especially to block IgG receptors on the cell surface to reduce nonspecific background.57 Generally, the serum for blocking IgG receptors should be from a species unrelated to the primary antibody and preferably from the same species as the secondary antibody.57
In each of the present protocols, the choice of primary antibody is the most critical step in achieving successful immunostaining. The antibody with the greatest specificity, purity, affinity, and avidity is desired. A good starting concentration for monoclonal antibodies is usually 1-5 µg/ml. Typical dilutions of the stock preparation of polyclonal antibodies range from 1:20 to 1:500.57 One needs to determine empirically the appropriate concentration of the primary antibody and the diluent for each sample. Samples should be gently rocked but not dried during incubation for homogenous staining. Extensive washing between the incubation of primary antibody and secondary antibody may be required to avoid cross reaction with immunoglobulins from other species. However, the optimal washing condition must be determined empirically to avoid the loss of cells, especially mitotic cells, which round off and detach when shaken (i.e., mitotic shake-off).58 For success, a secondary antibody must be chosen to bind to the primary antibody with high affinity. For instance, to avoid nonspecific binding of the Fc portion of the secondary antibody to IgG Fc receptors, Fb(ab')2 fragments can be used instead of the whole antibody.57 For many secondary antibodies, incubation time can be minimized to 30 min at RT to reduce the nonspecific background.
In addition to Protocol 5, laser-scanning confocal microscopy could be advantageous in detecting and analyzing the localization and the function of centromere-kinetochore proteins with the fluorescent properties. As shown in the List of Materials/Equipment, Alexa Fluor 488 and Alexa Fluor 594 are probably the most frequently used fluorescent dyes in the present protocols. To detect two and/or multiple different centromere-kinetochore proteins in the sample, one must ensure that the antibodies used to detect one protein do not prevent the detection of the second protein.57 In the present protocols, two primary antibodies are mixed and applied at the same time. However, one should optimize the immunostaining conditions for each protein separately before applying the primary or secondary antibodies together: if the two proteins are in close proximity as in the case of two centromere-kinetochore components, one primary antibody may structurally prevent the binding of the second primary antibody. Another concern of multiple signal detection is the problem of spectral overlap: the difficulty in preventing signal from one dye "leaking" into the spectral channel of another. This problem becomes more difficult for conventional interference filter sets to solve as the number of dyes increases or if the sample contains overenhanced signals (e.g., ACA or anti-Flag signals in the present study), including the interference of autofluorescence. Troubleshooting of autofluorescence has been conducted in other studies.57,59,60 In the present study, optimizing fluorescent filter sets with single-label controls in each channel is sufficient in most cases to avoid these problems (see below).
In each of the present protocols, proper controls are essential. Every new primary antibody should be characterized before immunostaining begins. The specificity of the antibody should be confirmed by Western blot analysis, if applicable. In addition, confirmation that the staining is not caused by nonspecific binding to the cell surface should be obtained, and the optimal working concentration of a particular primary antibody should be determined through the use of serial dilutions (i.e., a binding curve should be developed). A negative control (i.e., the absence of the primary antibody from the staining process) should be included. Another option of negative control is the substitution of normal IgG from the same species for the primary antibody. For detection of multiple labels, one must prepare controls without secondary antibodies (the background control) as well as single-label controls to avoid spectral overlap artifacts (i.e., bleed-through, crossover, crosstalk, or dye "leaking," as described above). One can quantify the fraction of "leaking" by comparing signals through the "wrong" filters with the correct signal, using these proportions to remove computationally all "leaking," and computing the real distribution and/or co-localization of fluorescent label.60 The background control should be examined independently with each channel to set the limits of signal gain and offset to be adapted for the final imaging. All channels that will be used to obtain an image of a multiple-label sample must be subjected to independent background correction, because the level of autofluorescence in each channel varies substantially. The "leaking" controls as described above are required to determine the amount of signal gain possible in each channel without detecting "leaking" into adjacent channels.60
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was supported by NIH grant GM68418.
Materials
| Lipofectamin 2000 | Life Technologies/Invitrogen | 11668 | transfection reagent I |
| Lipofectamin RNAiMAX | Life Technologies/Invitrogen | 13778 | transfection reagent II |
| Opti-MEM I | Life Technologies/Invitrogen | 31985 | Reduced serum media, warm in 37 °C water bath before use |
| High-glucose DMEM (Dulbecco’s modified Eagle’s medium) | Life Technologies/BioWhittaker | 12-604 | high-glucose DMEM, warm in 37 °C water bath before use |
| Fetal Bovine Serum, certified, heat inactivated, US origin | Life Technologies/Gibco | 10082 | FBS (fetal bovine serum) |
| Poly-L-Lysine SOLUTION | SIGMA-SLDRICH | P 8920 | Poly-L-Lysine, 0.1% w/v, in water |
| UltraPure Distilled Water | Life Technologies/Invitrogen/Gibco | 10977 | Sterile tissue culture grade water |
| Micro Cover glass (22 mm x 22 mm) | Surgipath | 105 | Cover glass (22 mm x 22 mm) |
| 6 Well Cell Culture Cluster | Fisher/Corning Incorporated | 07-200-83 | 6-well polystyrene plate |
| Penicillin, Streptomycin; Liquid | Fisher/Gibco | 15-140 | Penicillin-streptomycin |
| PAP PEN | Binding Site | AD100.1 | Hydrophobic barrier pen (for a water repellant barrier in immunofluorescent staining) |
| Paclitaxel (Taxol) | SIGMA-SLDRICH | T7402 | Taxol for mitotic cell analysis |
| TN-16, microtubule inhibitor (TN16) | Enzo Life Sciences | BML-T120 | TN16 for mitotic cell analysis |
| BSA (bovine serum albumin) | SIGMA-SLDRICH | A7906 | Blocking reagent |
| Triton X-100 | SIGMA-SLDRICH | T8787 | Detergent for permeabilization |
| Paraformaldehyde | SIGMA-SLDRICH | P6148 | Fixation reagant |
| DAPI | SIGMA-SLDRICH | D9542 | For nuclear staining |
| p-phenylenediamine | SIGMA-SLDRICH | P6001 | For mounting medium |
| VWR Micro Slides, Frosted | VWR International | 48312-013 | Micro slides |
| Anti-CENP-A antibody | Stressgen/Enzo Life Sciences | KAM-CC006 | Mouse monoclonal antibody; dilution ratio of 1:100 (IIF), 1:5000 (WB) |
| Anti-CENP-B antibody | Novus Biologicals | H00001059-B01P | Mouse monoclonal antibody; dilution ratio of 1:200 (IIF, methanol/acetone fixation)-1:400 (IIF, paraformaldehyde fixation) |
| Anti-CENP-B antibody | abcam | ab25734 | Rabbit polyclonal antibody; dilution ratio of 1:200 (IIF, methanol/acetone fixation)-1:400 (IIF, paraformaldehyde fixation) |
| Anti-centromere antibody (ACA) | Fitzgerald Industries International, Inc. | 90C-CS1058 | Human centromere antiserum; dilution ratio of 1:2000 (IIF) |
| Anti-CENP-H antibody | Bethyl Laboratories | BL1112 (A400-007A) | Rabbit polyclonal antibody; dilution ratio of 1:200 (IIF) |
| Anti-CENP-H antibody | BD | 612142 | Mouse monoclonal antibody; dilution ratio of 1:200 (IIF) |
| Anti-CENP-I antibody | N/A, Dr. Katusmi Kitagawa | N/A, Dr. Katusmi Kitagawa | Rabbit polyclonal antibody; dilution ratio of 1:1000 (IIF); Niikura et al., Oncogene, 4133-4146 (2006) |
| Anti-KNL1 antibody | Novus Biologicals | NBP1-89223 | Rabbit polyclonal antibody; dilution ratio of 1:100 (IIF) |
| Anti-Hec1 antibody | Novus Biologicals / GeneTex | NB 100-338 / GTX70268 | Mouse monoclonal antibody; dilution ratio of 1:100 (IIF) |
| Anti-Hec1 antibody | GeneTex | GTX110735 | Rabbit polyclonal antibody; dilution ratio of 1:100 (IIF) |
| Anti-Ska1 antibody | abcam | ab46826 | Rabbit polyclonal antibody; dilution ratio of 1:100 (IIF) |
| Anti-Flag antibody | SIGMA-ALDRICH | F3165 | Mouse monoclonal antibody; dilution ratio of 1:1000 (IIF), 1:5000 (WB) |
| Anti-Flag antibody | SIGMA-ALDRICH | F7425 | Rabbit polyclonal antibody; dilution ratio of 1:1000 (IIF), 1:5000 (WB) |
| Anti-CUL4A antibody | N/A, Dr. Pradip Raychaudhuri | N/A, Dr. Pradip Raychaudhuri | Rabbit polyclonal antibody; dilution ratio of 1:3000 (WB); Shiyanov et al., The Journal of biological chemistry, 35309-35312 (1999) |
| Anti-RBX1 antibody | Cell Signaling | 4397 | Rabbit polyclonal antibody; dilution ratio of 1:2000 (WB) |
| Anti-GAPDH antibody | Chemicon | MAB374 | Mouse monoclonal antibody; dilution ratio of 1:5000 (WB) |
| Alexa Fluor 488 Goat Anti-Mouse IgG | Life Technologies/Invitrogen | A11001 | fluorophore-conjugated secondary antibody (Affinity-purified secondary antibody) |
| Alexa Fluor 594 Goat Anti-Mouse IgG | Life Technologies/Invitrogen | A11005 | fluorophore-conjugated secondary antibody (Affinity-purified secondary antibody) |
| Alexa Fluor 488 Goat Anti-Rabbit IgG | Life Technologies/Invitrogen | A11008 | fluorophore-conjugated secondary antibody (Affinity-purified secondary antibody) |
| Alexa Fluor 594 Goat Anti-Rabbit IgG | Life Technologies/Invitrogen | A11012 | fluorophore-conjugated secondary antibody (Affinity-purified secondary antibody) |
| Non fat powdered milk (approved substitution for carnation powdered milk) | Fisher Scientific | NC9255871 (Reorder No. 190915; Lot# 90629) | Skim milk |
| Leica DM IRE2 motorized fluorescence microscope | Leica | motorized fluorescence microscope | |
| HCX PL APO 63x oil immersion lens | Leica | LEICA HCX PL APO NA 1.40 OIL PH 3 CS | 63X oil immersion lens |
| HCX PL APO 100x oil immersion lens | Leica | LEICA HCX PL APO NA 1.40 OIL PHE | 100X oil immersion lens |
| Leica EL6000 compact light source | Leica | External compact light source for fluorescent excitation | |
| ORCA-R2 Digital CCD camera | Hamamatsu | C10600-10B | digital CCD camera |
| Openlab version 5.5.2 Scientific Imaging Software | Perkin Elmer/Improvision | For image observation, acquisition, quantification, and analysis | |
| Velocity version 6.1.1 3D Image Analysis Software | Perkin Elmer/Improvision | For image observation, acquisition, quantification, and analysis | |
| Complete EDTA-free protease inhibitor cocktail | Roche | 11873580001/11836170001 | Protease inhibitor cocktail tablets |
| PlusOne 2-D Quant Kit | Amersham Biosciences | 80-6483-56 | Commercial protein assay reagent I for measurement of protein concentration (compatible with 2% SDS) |
| Bio-Rad Protein Assay | Bio-Rad | 500-0006 | Commercial protein assay reagent II for measurement of protein concentration (compatible with 0.1% SDS) |
| Immobilon-FL | EMD Millipore | IPFL00010 | PVDF membrane for transferring |
| IRDye 800CW Goat Anti-Mouse IgG | LI-COR Biosciences | 926-32210 | IR fluorescent dye-conjugated secondary antibody (Affinity-purified secondary antibody); dilution ratio of 1:20000 (IIF) |
| IRDye 680 Goat Anti-Rabbit IgG | LI-COR Biosciences | 926-32221 | IR fluorescent dye-conjugated secondary antibody (Affinity-purified secondary antibody); dilution ratio of 1:20000 (IIF) |
| Goat anti-Mouse IgG DyLight 549 | Fisher Scientific | PI35507 | DyLight-conjugated secondary antibodyIR fluorescent dye-conjugated secondary antibody (Affinity-purified secondary antibody); dilution ratio of 1:20000 (IIF) |
| Goat anti-Rabbit DyLight 649 | Fisher Scientific | PI35565 | DyLight-conjugated secondary antibodyIR fluorescent dye-conjugated secondary antibody (Affinity-purified secondary antibody); dilution ratio of 1:20000 (IIF) |
| Goat anti-mouse IgG-HRP | Santa Cruz | SC-2005 | HRP-conjugated secondary antibodyDyLight-conjugated secondary antibodyIR fluorescent dye-conjugated secondary antibody (Affinity-purified secondary antibody); dilution ratio of 1:10000 (IIF) |
| Goat anti-rabbit IgG-HRP | Santa Cruz | SC-2004 | HRP-conjugated secondary antibodyDyLight-conjugated secondary antibodyIR fluorescent dye-conjugated secondary antibody (Affinity-purified secondary antibody); dilution ratio of 1:10000 (IIF) |
| Openlab version 5.5.2. Scientific Imaging Software | Improvision/PerkinElmer | Software A | |
| Volocity version 6.3 3D Image Analysis Software (Volocity Acquisition) | PerkinElmer | Software B1 | |
| Volocity version 6.3 3D Image Analysis Software (Volocity Quantification) | PerkinElmer | Software B2 | |
| Branson SONIFIER 450 | Sonicator | ||
| Branson Ultrasonics sonicator Microtip Step, Solid, Threaded 9.5 mm | VWR Scientific Products Inc. | 33995-325 | Disruptor horn for sonication |
| Branson Ultrasonics sonicator Microtip Tapered 6.5 mm | VWR Scientific Products Inc. | 33996-185 | Microtip for sonication |
| Odyssey CLx Infrared imaging System | LI-COR Biosciences | Infrared imaging system for immunoblot detection | |
| Image Studio Analysis Software Ver 4.0 | LI-COR Biosciences | Software C | |
| Molecular Imager Versadoc MP4000 System | Bio-Rad | Chemiluminescence imager for immunoblot detection | |
| Quantity One 1-D analysis software | Bio-Rad | Software D | |
| SuperSignal West Femto Maximum Sensitivity Substrate | Thermo | 34095 | Ultra-sensitive enhanced chemiluminescent (ECL) substrate |
References
- DeLuca, J. G., et al. Hec1 and nuf2 are core components of the kinetochore outer plate essential for organizing microtubule attachment sites. Molecular biology of the cell. 16, 519-531 (2005).
- Earnshaw, W. C., Halligan, N., Cooke, C., Rothfield, N. The kinetochore is part of the metaphase chromosome scaffold. J Cell Biol. 98, 352-357 (1984).
- Hoffman, D. B., Pearson, C. G., Yen, T. J., Howell, B. J., Salmon, E. D. Microtubule-dependent changes in assembly of microtubule motor proteins and mitotic spindle checkpoint proteins at PtK1 kinetochores. Molecular biology of the cell. 12, 1995-2009 (2001).
- Regnier, V., et al. CENP-A is required for accurate chromosome segregation and sustained kinetochore association of BubR1. Mol Cell Biol. 25, 3967-3981 (2005).
- Bernad, R., Sanchez, P., Losada, A. Epigenetic specification of centromeres by CENP-A. Exp Cell Res. 315, 3233-3241 (2009).
- Black, B. E., Cleveland, D. W. Epigenetic centromere propagation and the nature of CENP-a nucleosomes. Cell. 144, 471-479 (2011).
- Warburton, P. E., et al. Immunolocalization of CENP-A suggests a distinct nucleosome structure at the inner kinetochore plate of active centromeres. Curr Biol. 7, 901-904 (1997).
- Karpen, G. H., Allshire, R. C. The case for epigenetic effects on centromere identity and function. Trends Genet. 13, 489-496 (1997).
- Black, B. E., et al. Structural determinants for generating centromeric chromatin. Nature. 430, 578-582 (2004).
- Black, B. E., et al. Centromere identity maintained by nucleosomes assembled with histone H3 containing the CENP-A targeting domain. Mol Cell. 25, 309-322 (2007).
- Fachinetti, D., et al. A two-step mechanism for epigenetic specification of centromere identity and function. Nat Cell Biol. 15, 1056-1066 (2013).
- Zeitlin, S. G., Shelby, R. D., Sullivan, K. F. CENP-A is phosphorylated by Aurora B kinase and plays an unexpected role in completion of cytokinesis. The Journal of cell biology. 155, 1147-1157 (2001).
- Zhang, X., Li, X., Marshall, J. B., Zhong, C. X., Dawe, R. K. Phosphoserines on maize CENTROMERIC HISTONE H3 and histone H3 demarcate the centromere and pericentromere during chromosome segregation. The Plant cell. 17, 572-583 (2005).
- Goutte-Gattat, D., et al. Phosphorylation of the CENP-A amino-terminus in mitotic centromeric chromatin is required for kinetochore function. Proc Natl Acad Sci U S A. 110, 8579-8584 (2013).
- Bailey, A. O., et al. Posttranslational modification of CENP-A influences the conformation of centromeric chromatin. Proc Natl Acad Sci U S A. 110, 11827-11832 (2013).
- Samel, A., Cuomo, A., Bonaldi, T., Ehrenhofer-Murray, A. E. Methylation of CenH3 arginine 37 regulates kinetochore integrity and chromosome segregation. Proc Natl Acad Sci U S A. 109, 9029-9034 (2012).
- Niikura, Y., et al. CENP-A K124 Ubiquitylation Is Required for CENP-A Deposition at the Centromere. Dev Cell. , (2015).
- Chan, G. K., Liu, S. T., Yen, T. J. Kinetochore structure and function. Trends in cell biology. 15, 589-598 (2005).
- Hori, T., Okada, M., Maenaka, K., Fukagawa, T. CENP-O class proteins form a stable complex and are required for proper kinetochore function. Molecular biology of the cell. 19, 843-854 (2008).
- Fukagawa, T., Earnshaw, W. C. The centromere: chromatin foundation for the kinetochore machinery. Developmental cell. 30, 496-508 (2014).
- Kedersha, N., Grainger, D. . The Proteintech Blog.Proteintech. , (2012).
- Clontech Laboratories, Inc. . HeLa Tet-Off Advanced Cell Line. , (2012).
- Meraldi, P., Sorger, P. K. A dual role for Bub1 in the spindle checkpoint and chromosome congression. EMBO J. 24, 1621-1633 (2005).
- Niikura, Y., et al. 17-AAG, an Hsp90 inhibitor, causes kinetochore defects: a novel mechanism by which 17-AAG inhibits cell proliferation. Oncogene. 25, 4133-4146 (2006).
- Yang, Z., et al. Silencing mitosin induces misaligned chromosomes, premature chromosome decondensation before anaphase onset, and mitotic cell death. Mol Cell Biol. 25, 4062-4074 (2005).
- Wang, H., et al. Histone H3 and H4 ubiquitylation by the CUL4-DDB-ROC1 ubiquitin ligase facilitates cellular response to DNA damage. Mol Cell. 22, 383-394 (2006).
- Lamb, J. R., Tugendreich, S., Hieter, P. Tetratrico peptide repeat interactions: to TPR or not to TPR?. Trends Biochem Sci. 20, 257-259 (1995).
- Kitagawa, K., Skowyra, D., Elledge, S. J., Harper, J. W., Hieter, P. SGT1 encodes an essential component of the yeast kinetochore assembly pathway and a novel subunit of the SCF ubiquitin ligase complex. Mol Cell. 4, 21-33 (1999).
- Niikura, Y., Dixit, A., Scott, R., Perkins, G., Kitagawa, K. BUB1 mediation of caspase-independent mitotic death determines cell fate. J Cell Biol. 178, 283-296 (2007).
- Niikura, Y., Kitagawa, K. Identification of a novel splice variant: human SGT1B (SUGT1B). DNA Seq. 14, 436-441 (2003).
- Niikura, Y., Ogi, H., Kikuchi, K., Kitagawa, K. BUB3 that dissociates from BUB1 activates caspase-independent mitotic death (CIMD). Cell Death Differ. 17, 1011-1024 (2010).
- Ando, S., Yang, H., Nozaki, N., Okazaki, T., Yoda, K. CENP-A, -B, and -C chromatin complex that contains the I-type alpha-satellite array constitutes the prekinetochore in HeLa cells. Mol Cell Biol. 22, 2229-2241 (2002).
- Izuta, H., et al. Comprehensive analysis of the ICEN (Interphase Centromere Complex) components enriched in the CENP-A chromatin of human cells. Genes to cells : devoted to molecular & cellular mechanisms. 11, 673-684 (2006).
- Obuse, C., et al. Proteomics analysis of the centromere complex from HeLa interphase cells: UV-damaged DNA binding protein 1 (DDB-1) is a component of the CEN-complex, while BMI-1 is transiently co-localized with the centromeric region in interphase. Genes Cells. 9, 105-120 (2004).
- Merlet, J., Burger, J., Gomes, J. E., Pintard, L. Regulation of cullin-RING E3 ubiquitin-ligases by neddylation and dimerization. Cell Mol Life Sci. 66, 1924-1938 (2009).
- Bennett, E. J., Rush, J., Gygi, S. P., Harper, J. W. Dynamics of cullin-RING ubiquitin ligase network revealed by systematic quantitative proteomics. Cell. 143, 951-965 (2010).
- Antonelli, A., et al. Efficient inhibition of macrophage TNF-alpha production upon targeted delivery of K48R ubiquitin. Br J Haematol. 104, 475-481 (1999).
- Codomo, C. A., Furuyama, T., Henikoff, S. CENP-A octamers do not confer a reduction in nucleosome height by AFM. Nat Struct Mol Biol. 21, 4-5 (2014).
- Thrower, J. S., Hoffman, L., Rechsteiner, M., Pickart, C. M. Recognition of the polyubiquitin proteolytic signal. The EMBO journal. 19, 94-102 (2000).
- Yoda, K., et al. Human centromere protein A (CENP-A) can replace histone H3 in nucleosome reconstitution in vitro. Proc Natl Acad Sci U S A. 97, 7266-7271 (2000).
- Shelby, R. D., Vafa, O., Sullivan, K. F. Assembly of CENP-A into centromeric chromatin requires a cooperative array of nucleosomal DNA contact sites. J Cell Biol. 136, 501-513 (1997).
- Majumder, S., Fisk, H. A. Quantitative immunofluorescence assay to measure the variation in protein levels at centrosomes. J Vis Exp. , (2014).
- Masumoto, H., Masukata, H., Muro, Y., Nozaki, N., Okazaki, T. A human centromere antigen (CENP-B) interacts with a short specific sequence in alphoid DNA, a human centromeric satellite. J Cell Biol. 109, 1963-1973 (1989).
- Yoda, K., Kitagawa, K., Masumoto, H., Muro, Y., Okazaki, T. A human centromere protein, CENP-B, has a DNA binding domain containing four potential alpha helices at the NH2 terminus, which is separable from dimerizing activity. J Cell Biol. 119, 1413-1427 (1992).
- Sugata, N., et al. Human CENP-H multimers colocalize with CENP-A and CENP-C at active centromere–kinetochore complexes. Hum Mol Genet. 9, 2919-2926 (2000).
- Earnshaw, W. C., Ratrie, H., Stetten, G. Visualization of centromere proteins CENP-B and CENP-C on a stable dicentric chromosome in cytological spreads. Chromosoma. 98, 1-12 (1989).
- Goshima, G., Kiyomitsu, T., Yoda, K., Yanagida, M. Human centromere chromatin protein hMis12, essential for equal segregation, is independent of CENP-A loading pathway. J Cell Biol. 160, 25-39 (2003).
- Liu, S. T., Rattner, J. B., Jablonski, S. A., Yen, T. J. Mapping the assembly pathways that specify formation of the trilaminar kinetochore plates in human cells. J Cell Biol. 175, 41-53 (2006).
- Gascoigne, K. E., Cheeseman, I. M. T time for point centromeres. Nat Cell Biol. 14, 559-561 (2012).
- Gascoigne, K. E., et al. Induced ectopic kinetochore assembly bypasses the requirement for CENP-A nucleosomes. Cell. 145, 410-422 (2011).
- Nishino, T., et al. CENP-T provides a structural platform for outer kinetochore assembly. EMBO J. 32, 424-436 (2013).
- Malvezzi, F., et al. A structural basis for kinetochore recruitment of the Ndc80 complex via two distinct centromere receptors. EMBO J. 32, 409-423 (2013).
- Schleiffer, A., et al. CENP-T proteins are conserved centromere receptors of the Ndc80 complex. Nat Cell Biol. 14, 604-613 (2012).
- Rago, F., Gascoigne, K. E., Cheeseman, I. M. Distinct organization and regulation of the outer kinetochore KMN network downstream of CENP-C and CENP-T. Curr Biol. 25, 671-677 (2015).
- Nishino, T., et al. CENP-T-W-S-X forms a unique centromeric chromatin structure with a histone-like fold. Cell. 148, 487-501 (2012).
- Bodor, D. L., et al. The quantitative architecture of centromeric chromatin. Elife. 3, e02137 (2014).
- Oliver, C., Rapely, R., Walker, J. M. Ch 58. Molecular Biomethods Handbook. , 1063-1079 (2008).
- Terasima, T., Tolmach, L. J. Changes in x-ray sensitivity of HeLa cells during the division cycle. Nature. 190, 1210-1211 (1961).
- Levenson, G. B. R., Vo-Dinh, T. u. a. n. Ch 8. Biomedical Photonics Handbook. , 8-19 (2003).
- Sanderson, J. . Fluorescence bleed-though. , (2011).

