Growth Curves: Generating Growth Curves Using Colony Forming Units and Optical Density Measurements
English
Share
Overview
Source: Andrew J. Van Alst1, Rhiannon M. LeVeque1, Natalia Martin1, and Victor J. DiRita1
1 Department of Microbiology and Molecular Genetics, Michigan State University, East Lansing, Michigan, United States of America
Growth curves provide valuable information on bacterial growth kinetics and cell physiology. They allow us to determine how bacteria respond in variable growth conditions as well as to define optimal growth parameters for a given bacterium. An archetypal growth curve progresses through four stages of growth: lag, exponential, stationary, and death (1).
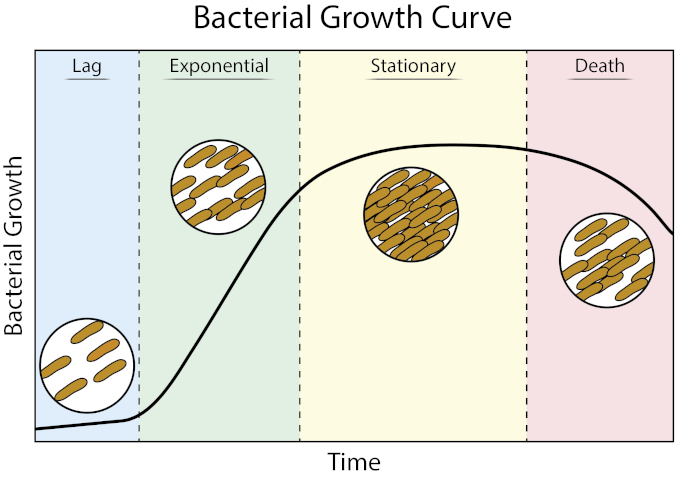
Figure 1: Bacterial growth curve. Bacteria grown in batch culture progress through four phases of growth: lag, exponential, stationary, and death. Lag phase is the period of time it takes for the bacteria to reach a physiological state capable of rapid cell growth and division. Exponential phase is the stage of fastest cell growth and division during which DNA replication, RNA transcription, and protein production all occur at a constant, rapid rate. Stationary phase is characterized by a slowing down and plateauing of bacterial growth due to nutrient limitation and/or toxic intermediate accumulation. Death phase is the stage during which cell lysis occurs as a result of severe nutrient limitation.
Lag phase is the period of time it takes for the bacteria to reach a physiological state capable of rapid cell growth and division. This lag occurs because it takes time for bacteria to adjust to their new environment. Once the necessary cellular components are generated in lag phase, bacteria enter the exponential phase of growth where DNA replication, RNA transcription, and protein production all occur at a constant, rapid rate (2). The rate of rapid cell growth and division during the exponential phase is calculated as the generation time, or doubling time, and is the fastest rate at which the bacteria can replicate under the given conditions (1). The doubling time can be used to compare different growth conditions to determine which is more favorable for bacterial growth. The exponential growth phase is the most reproducible growth condition as bacterial cell physiology is consistent throughout the entire population (3). Stationary phase follows the exponential phase where cell growth plateaus. Stationary phase is brought on due to nutrient depletion and/or accumulation of toxic intermediates. Bacterial cells continue to survive in this stage, although the rate of replication and cell division is drastically reduced. The final phase is death, where severe nutrient depletion leads to the lysing of cells. Features of the growth curve that provide the most information include the duration of lag phase, the doubling time, and the maximum cell density reached.
Quantification of bacteria in batch culture can be determined using both colony forming units and optical density measurements. Enumeration by colony forming units (CFU) provides a direct measurement of bacterial cell counts. The standard unit of measure for CFU is the number of culturable bacteria present per 1 mL of culture (CFU/mL) determined by serial dilution and spread plating techniques. For each timepoint, a 1:10 dilution series of the batch culture is performed and 100 µl of each dilution is spread plated using a cell spreader.
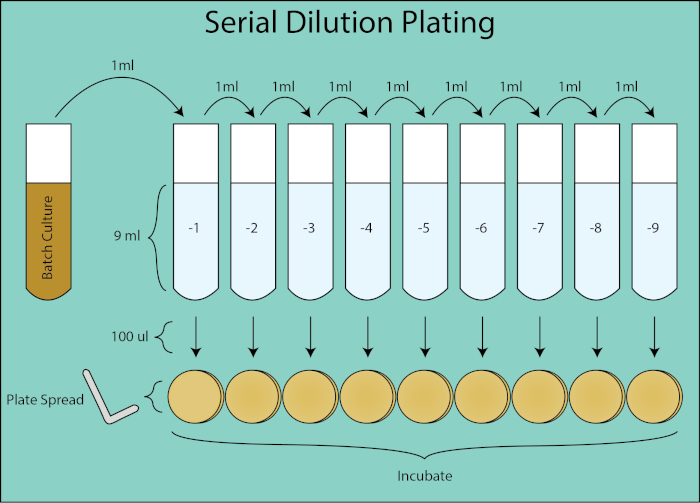
Figure 2. Serial dilution plating schematic. General flowthrough for dilution plating from batch culture. The batch culture is serially diluted 1:10 by transferring 1 mL of the previous dilution into the subsequent tube containing 9ml PBS. From each dilution tube, 100 µl is spread plated using a plate spreader which is an additional dilution of 1:10 as it is 1/10th the volume of 1 mL volume when calculating CFU/mL. Plates are incubated and enumerated once clonal colonies grow on the plates.
The plates are then incubated overnight and clonal colonies enumerated. The dilution plate which grew 30-300 colonies is used to calculate the CFU/mL for the given timepoint (4, 5). Stochastic variation in colony counts under 30 are subject to greater error in the calculation of CFU/mL and counting colonies greater than 300 can be underestimated due to colony crowding and overlapping. Using the dilution factor for the given plate, the CFU of the batch culture can be calculated for each timepoint.
Optical density gives an instant approximation of bacterial cell count measured using a spectrophotometer. The optical density is a measure of absorbance of light particles that pass through 1cm of culture and detected by a photodiode sensor (6). The optical density of a culture is measured in relation to a media blank and increases as bacterial density increases. For bacterial cells, a wavelength of 600 nm (OD600) is typically used when measuring optical density (4). By generating a standard curve relating colony forming units and optical density, the optical density measurement can be used to readily approximate the bacterial cell count of a batch culture. However, this relationship begins to deteriorate as early as 0.3 OD600 as cells begins to change shape and accumulate extracellular products in the media, influencing the optical density reading as it relates to CFU (7). This error becomes more pronounced during stationary and death phases.
Here, Escherichia coli is grown in Luria-Bertani (LB) broth at 37°C over the course of 30 hours (7). Both CFU/mL and optical density growth curves have been generated as well as the standard curve relating optical density to CFU.
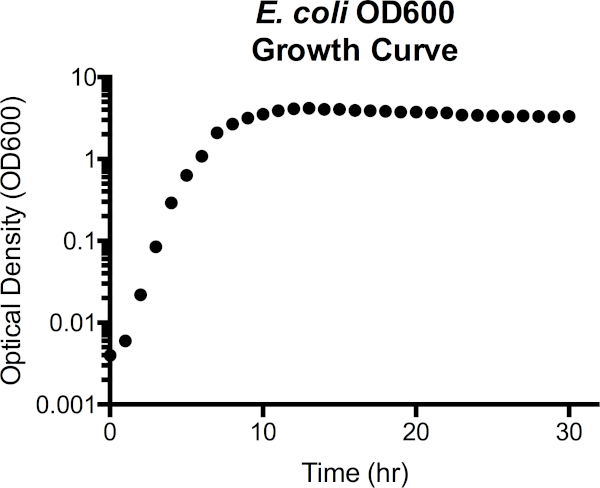
Figure 3. Escherichia coli optical density at 600 nm wavelength (OD600) growth curve. Optical density values were taken directly from the spectrophotometer after blanking with sterile LB media. OD600 values greater than 1.0 were diluted 1:10 by combining 100 µl culture with 900 µl fresh LB, again measured, and then multiplied by 10 to obtain the OD600 value. This step is taken as the accuracy in measurement of the spectrophotometer is reduced at high cell density. From the curve, lag phase extends to around 1h of growth, transitions to exponential phase from 2h to 7h, then begins to plateau, entering stationary phase. Death phase is not a stark transition, however, as optical density gradually begins to decline after 15h.

Figure 4. Escherichia coli colony forming unit per milliliter (CFU/mL) growth curve. CFU/mL values for each timepoint were calculated from the dilution plate that contained 30-300 colonies. From the curve, lag phase extends out to around 2h of growth, transitions to exponential phase from 2h to 7h, then begins to plateau, entering stationary phase. Death phase is not a stark transition, however, as CFU/mL gradually begins to decline after 15h from a peak of 2 x 109 to approximately 5 x 108 at 30 hours.
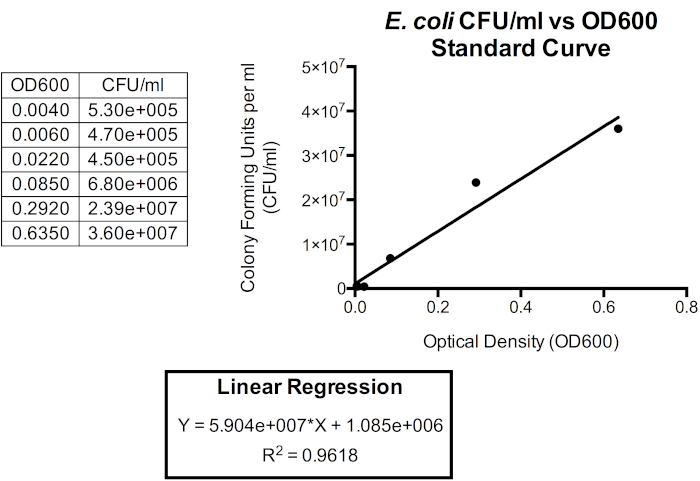
Figure 5. Standardization curve for CFU/mL versus OD600. A linear regression can be used to relate these units so that optical density may be used to approximate bacterial cell density. Optical density can be used to provide and instant approximation of the CFU/mL of the batch culture. Here, only the first six timepoints are plotted as the relationship between OD600 and CFU/mL is less accurate beyond 1.0 OD600 as cell shape and extracellular products begin to accumulate as the bacteria enter stationary phase, which occurs shortly after reaching 1.0 OD600. Changes in cell shape and extracellular products in the media influence the optical density reading and therefore the relationship between optical density and the number of bacteria in the culture is also impacted.
The doubling time has also been determined to be 15 minutes and 19 seconds. From this data, the capacity for growth in LB for E. coli can be visualized and be used for comparison between different media or bacteria.
Procedure
1. Set-up
- Required laboratory materials: liquid media, solidified agar media, Erlenmeyer flasks, 15 mL test tubes, phosphate buffered saline (PBS), bacterial cell spreader, 70% ethanol, and a spectrophotometer. All solutions and glassware must be sterilized prior to use.
- Prepare the work station by sterilizing with 70% ethanol. Work near a Bunsen burner to prevent contamination of media.
- When working with bacteria, proper personal protective equipment and aseptic technique should be used. A lab coat and gloves are required when working with bacterial cultures.
- Recipes for buffers, solutions, and reagents
- Phosphate buffered saline (PBS) (8).
- Luria-Bertani Broth (LB) (9).
2. Protocol
- Preparation of Media
- Identify the growth media with which to grow the bacteria and prepare both liquid broth and solid agar (1.5% w/v agar) media in separate autoclavable bottles. Here, LB broth and LB agar were prepared for the growth of Escherichia coli.
- Sterilize the media with a semi-tightened cap in an autoclave set to 121 °C for 35 min.
- For agar media, after autoclaving, place in a water bath set to 50 °C for 30 minutes to cool. Once cooled, pour 20-25 mL agar media into 100x15mm circular Petri dishes. Allow plates to set 24 hours at room temperature before use.
- Initial Preparation of Bacteria
- From frozen stock, streak bacteria for isolation on selected media agar to obtain single colony isolates. Incubate in growth conditions permissible for the chosen bacteria. Here, E. coli is streaked on LB agar and is incubated at 37 °C overnight (16-18h).
- Using a sterile inoculation loop, select a single colony from the streak plate and inoculate 4 mL liquid media in a 15 mL test tube and grow in conditions permissible for the chosen bacteria. Here, E. coli is grown at 37 °C with shaking at 210 rpm overnight (16-18h).
- Growth Curve Setup
- Growth flask preparation
- Autoclave appropriately sized Erlenmeyer flasks. Typically, a 1:5 ratio of media to total flask volume is used. Here, 100 mL LB media is used in a 500 mL flask.
- Using a serological pipette, transfer sterile media to the Erlenmeyer flask.
- Dilution series preparation
- Label 15 mL test tubes: -1, -2, -3, -4, -5, -6, -7, -8, and -9, distributing 9mL PBS into each. These numbers correspond to the dilution factor used to calculate CFU/mL. A new set of tubes is needed for each collection timepoint. (Figure 2)
- Agar plate preparation
- Label plates with time of collection and dilution factor. For each timepoint there will be one plate for each dilution.
- Growth flask preparation
- Growth Curve Protocol
- Inoculation of media
- Using the overnight liquid culture prepared as part of step 2.2.2, inoculate the flask media with 1:1000 volume of culture. Here, 100 µL overnight liquid culture is added to 100 mL LB media.
- Swirl the media to evenly distribute the bacteria.
- Timepoint collection
- Growth condition setup
- Place flask in experimental growth conditions chosen for the given bacteria. Timepoints should be taken frequently for fast-growing bacteria and can be taken in longer intervals for slow-growing bacteria. Here, E. coli is grown at 37°C with shaking at 210 revolutions per minute (rpm) and timepoints taken every 1 hour.
- Optical density (OD600) measurement
- At each timepoint, including the starting timepoint (t = 0), withdraw 1 mL of bacterial culture and dispense into a spectrophotometer cuvette.
- Wipe the cuvette clean and record the optical density at 600 nm wavelength. If the optical density reading is greater than 1.0, dilute 100 µL of culture 1:10 with 900 µL fresh media, record the optical density, and multiply this value by 10 for the OD600 measurement.
- Colony forming unit (CFU/mL) measurement
- At each timepoint, withdraw 1 mL of bacterial culture and dispense into the -1 glass test tube containing 9 mL of PBS.
- For the dilution series, serially transfer 1 mL from the -1 tube down all dilution tubes to the -9, vortexing after each transfer. (Figure 2)
- For each dilution, dispense 100 µL of cell suspension to the correspondingly labelled solid media agar plate. (Figure 2)
- Using a cell spreader that has been sterilized in ethanol, passed through a Bunsen burner flame, and cooled by touching the surface of the agar, spread the 100 µL of cell suspension until the surface of the agar plate becomes dry.
- Incubate the spread plates upside-down at a temperature that supports growth of the bacteria. Here, E. coli is incubated at 37°C.
- After incubation, once visible colonies arise, count the number of bacterial colonies on each plate and record these values along with their associated dilution factor for all plates at each timepoint.
- Growth condition setup
- Inoculation of media
3. Data Analysis and Results
- Optical Density (OD600) Growth Curve Plot
- Plot the optical density (OD600) versus time on a semi-log scale. (Figure 3)
- Colony Forming Unit (CFU/mL) Growth Curve Plot
- For each timepoint, choose the dilution plate where the colony counts fell within the range of 30-300 bacteria. Multiply the colony count number by the dilution factor and then by 10 as the 100 µL spread is considered an additional 1:10 dilution when calculating CFU/mL.

- Plot the colony forming units versus time on a semi-log scale. (Figure 4)
- For each timepoint, choose the dilution plate where the colony counts fell within the range of 30-300 bacteria. Multiply the colony count number by the dilution factor and then by 10 as the 100 µL spread is considered an additional 1:10 dilution when calculating CFU/mL.
- Relating optical density and colony forming units
- Plot the colony forming units versus optical density on a linear scale for OD600 readings less than or equal to 1.0 OD600 as the relationship between OD600 and CFU/mL is less accurate beyond 1.0 OD600. Here, the first six timepoints are plotted. (Figure 5)
- Generate a linear regression trendline displaying the equation and R2 value.
- Determining bacteria doubling time
- Using the colony forming unit growth curve plot, during exponential phase, identify two points on the graph with the steepest slope between them to calculate the doubling time.
- Calculating the doubling time

- ΔTime = t2 – t1, where t1 = Timepoint 1 and t2 = Timepoint 2
 , where b = number of bacteria at t2, B = number of bacteria at t1, and n = number of generations. Derived from:
, where b = number of bacteria at t2, B = number of bacteria at t1, and n = number of generations. Derived from: .
- Calculate doubling time using:

Bacteria reproduce through a process called cell division, which results in two identical daughter cells. If the growth conditions are favorable, bacterial populations will grow exponentially.
Bacterial growth curves plot the amount of bacteria in a culture as a function of time. A typical growth curve progresses through four stages: lag phase, exponential phase, stationary phase, and death phase. The lag phase is the time it takes for bacteria to reach a state where they can grow and divide quickly. After this, the bacteria transition to the exponential phase, characterized by rapid cell growth and division. The rate of exponential growth of the bacterial culture during this phase can be expressed as the doubling time, the fastest rate at which bacteria can reproduce under specific conditions. The stationary phase comes next, where bacterial cell growth plateaus and the growth and death rates even out due to environmental nutrient depletion. Finally, the bacteria enter the death phase. This is where bacterial growth declines sharply and severe nutrient depletion leads to the lysing of cells.
Two techniques can be used to quantify the amount of bacteria present in a culture and plot a growth curve. The first of these is via colony forming units, or CFUs. To obtain CFUs a one to ten series of nine dilutions is performed at regular time points. The first of these dilutions, negative one in this example, contains 9mL of PBS and 1mL of the bacterial culture. Resulting in a 1:10 dilution factor. Then, 1mL of this solution is transferred to the next tube, negative two, resulting in a 1:100 dilution factor. This process continues through the last tube, negative nine, resulting in a final dilution factor of 1:1 billion. After this, 100 microliters of each dilution is plated. The plates are then incubated and the clonal colonies are counted. The dilution plate for a given time point that grows between 30 and 300 colonies is used to calculate the CFUs per milliliter for that time point.
The second common method of measuring bacterial concentration is the optical density. The optical density of a culture can be measured instantly, in relation a media blank, with a spectrophotometer. Typically a wave length of 600 nanometers, also referred to as OD600, is used for these measurements, which increase as cell density increases. While optical density is less precise than CFUs, it is convenient because it can be obtained instantaneously and requires relatively few reagents. Both techniques can be used together to create a standard curve that more accurately approximates the bacterial cell count of a culture. In this video, you will learn how to obtain CFUs and OD600 measurements from timed serial dilutions of E. coli. Then, two growth curves using the CFU and OD600 measurements, respectively, will be plotted before being related by a standard curve.
When working with bacteria, it is important to use the appropriate personal protective equipment such as a lab coat and gloves and to observe proper aseptic technique.
After this, sterilize the work station with 70% ethanol. First, prepare the LB broth and LB solid agar media in separate autoclaveable bottles. After partially closing the caps of the bottles, sterilize the media in an autoclave set to 121 degrees Celsius for 35 minutes. Next, allow the agar media to cool in a water bath set to 50 degrees Celsius for 30 minutes. Once cooled, pour 20 to 25 mL into each Petri dish. After this, allow the plates to set for 24 hours at room temperature.
To prepare the single colony isolates that will later be used to produce a liquid bacterial culture, use previously frozen stock and proper streak plating technique to streak E. coli for isolation on LB agar. Incubate the dish at 37 degree Celsius overnight. After this, cool a flame sterilized inoculation loop on the agar before selecting a single colony from the streaked plate. Inoculate 4 mL of liquid media in a 15 mL test tube. Then, grow the E. coli at 37 degrees Celsius overnight with shaking at 210 rpm.
To set up the 1:1000 volume of bacterial culture that will be used in the growth curve, first obtain an autoclaved 500 mL Erlenmeyer flask. Then, use a 50 mL serological pipette to transfer 100 mL of sterile media to the flask. Next, label nine 15 ml test tubes consecutively as one through nine. These numbers correspond to the dilution factor that will be used to calculate the colony forming unit, or CFU. Then, add 9 mL of 1X PBS to each tube. After this, label the prepared agar plates with the corresponding time points and dilution factors that will be grown. In this example with E. coli, after the starting time point, time points are taken once every hour. Using the previously prepared overnight liquid E. coli culture, inoculate the media in the autoclave 500 mL Erlenmeyer flask with 1:1000 volume of culture. Swirl the media to evenly distribute the bacteria.
After blanking a spectrophotometer, clean the cuvette with a lint-free wipe. Next, dispense 1 mL of the culture into the cuvette and place it into the spectrophotometer to obtain the optical density of the culture at time point zero. Then, grow the E. coli at 37 degrees Celsius with shaking at 210 rpm. At each time point after time point zero, withdraw another 1 mL of bacterial culture from the flask and repeat the optical density measurement. If the optical density reading is greater than 1.0, dilute 100 microliters of bacterial culture with 900 microliters of fresh media and then measure the optical density once more. This value can be multiplied by 10 for the OD 600 measurement.
To obtain the colony forming unit measurement for each time point, withdraw an additional 1 mL of bacterial culture from the flask at each time point. Dispense the bacterial culture into the negative one test tube and vortex to mix. Then, perform the dilution series by first transferring 1 mL from the negative one tube into the negative two tube and vortex to mix. Transfer 1 mL from the negative two tube into the negative three tube and vortex to mix. Continue this serial transfer down all the dilution tubes to the negative nine tube. Dispense 100 microliters of cell suspension onto the correspondingly labeled plate for each dilution. For every dilution, sterilize a cell spreader in ethanol, pass it through a Bunsen burner flame, and cool it by touching the surface of the agar away from the inoculate. Then, use the cell spreader to spread the cell suspension until the surface of the agar plate becomes dry. Incubate the plates upside down at 37 degrees Celsius. Once visible colonies arise, count the number of bacterial colonies on each plate. Record these values and their associated dilution factors for each plate at each time point.
To create an OD 600 growth curve, after ensuring all the data points are entered correctly into a table, select all of the time points and their corresponding data. To generate a colony forming unit growth curve plot, choose the dilution plate where the colony counts fell within the range 30 to 300 bacteria for each time point. Multiply the colony count number by the dilution factor, and then by ten. This is because the 100 microliters spread is considered an additional 1:10 dilution when calculating colony forming units per milliliter. After this, plot the colony forming units versus time on a semi-log scale.
These plots produced with OD 600 and CFU measurements, respectively, can provide valuable information on E. coli growth kinetics. The optical density and colony forming units can be related, so that CFUs per milliliter can be estimated from OD 600 measurements, saving time and materials in future experiments.
To do this, plot the colony forming units against the optical density on a linear scale for OD 600 readings less than or equal to 1. 0. After this, generate a linear regression trend line in Y = MX + B format, where M is the slope and B is the y-intercept. Right click on the data points and select add trend line and linear. Then, check the box to display the equation on the chart and display the R squared value on the chart. The R squared value is the statistical measurement of how closely the data matched the fitted regression line. In this example, the first 6 time points are plotted with OD 600 on the x axis and CFUs per milliliter on the y axis. In future experiments with the same growth conditions, these slope and y-intercept values can be plugged into this equation to estimate CFUs from OD 600 readings. Next, look at the colony forming unit growth curve plot. During the exponential phase, identify two time points with the steepest slope between them. To calculate the doubling time, first calculate the change in time between the selected time points. Then, calculate the change in generations using the equation shown here. Here, lower case b is the number of bacteria at time point three and upper case B is the number of bacteria at time point two. Finally, divide the change in time by the change in generations. In this example, the doubling time is 0. 26 hours or 15 minutes and 19 seconds. Comparing doubling times across different experimental treatment allows us to identify the best growth conditions for a certain bacterial species. Therefore, the treatment with the lowest doubling time will be most optimal of the conditions tested.
Results
Plots of colony forming units and optical density are two ways to visualize growth kinetics. By determining the relationship between CFU/mL and OD600, the optical density plot also provides an estimate of CFU/mL over time. Conditions that result in the shortest doubling time are considered optimal for growth of the given bacteria.
Applications and Summary
Growth curves are valuable for understanding the growth kinetics and physiology of bacteria. They allow us to determine how bacteria respond in variable growth conditions as well as define the optimal growth parameters for a given bacterium. Colony forming unit and optical density plots both contain valuable information depicting the duration of lag phase, maximum cell density reached, and allowing for the calculation of bacterial doubling time. Growth curves also allow for comparison between different bacteria under the same growth conditions. Additionally, optical density provides a means of standardizing initial inoculums, enhancing consistency in other experiments.
Determining which approach to use when designing a growth curve experiment requires consideration. As the preferred method for generating growth curves, colony forming unit plots more accurately reflect the viable cell counts in batch culture. Colony forming unit plots also allow for measuring bacterial growth in conditions that would otherwise interfere with an optical density measurement. However, it is a more time consuming process, requiring extensive use of reagents, and must be performed manually. Optical density plots are less accurate and provide only an estimate of the colony forming units, requiring a standard curve to be generated for each unique bacteria. Optical density is primarily used for its convenience as it is far less time consuming and does not require many reagents to perform. What is most attractive to optical density, is that spectrophotometric incubators can automatically generate growth curves, vastly increasing the number of culture conditions that can be tested at once and eliminating the need to constantly attend the culture.
References
- R. E. Buchanan. 1918. Life Phases in a Bacterial Culture. J Infect Dis 23:109-125.
- CAMPBELL A. 1957. Synchronization of cell division. Bacteriol Rev 21:263-72.
- Wang P, Robert L, Pelletier J, Dang WL, Taddei F, Wright A, Jun S. 2010. Robust growth of Escherichia coli. Curr Biol 20:1099-103.
- Goldman E, Green LH. 2015. Practical Handbook of Microbiology, Third Edition. CRC Press.
- Ben-David A, Davidson CE. 2014. Estimation method for serial dilution experiments. J Microbiol Methods 107:214-221.
- Koch AL. 1968. Theory of the angular dependence of light scattered by bacteria and similar-sized biological objects. J Theor Biol 18:133-156.
- Sezonov G, Joseleau-Petit D, D'Ari R. 2007. Escherichia coli physiology in Luria-Bertani broth. J Bacteriol 189:8746-9.
Transcript
Bacteria reproduce through a process called cell division, which results in two identical daughter cells. If the growth conditions are favorable, bacterial populations will grow exponentially.
Bacterial growth curves plot the amount of bacteria in a culture as a function of time. A typical growth curve progresses through four stages: lag phase, exponential phase, stationary phase, and death phase. The lag phase is the time it takes for bacteria to reach a state where they can grow and divide quickly. After this, the bacteria transition to the exponential phase, characterized by rapid cell growth and division. The rate of exponential growth of the bacterial culture during this phase can be expressed as the doubling time, the fastest rate at which bacteria can reproduce under specific conditions. The stationary phase comes next, where bacterial cell growth plateaus and the growth and death rates even out due to environmental nutrient depletion. Finally, the bacteria enter the death phase. This is where bacterial growth declines sharply and severe nutrient depletion leads to the lysing of cells.
Two techniques can be used to quantify the amount of bacteria present in a culture and plot a growth curve. The first of these is via colony forming units, or CFUs. To obtain CFUs a one to ten series of nine dilutions is performed at regular time points. The first of these dilutions, negative one in this example, contains 9mL of PBS and 1mL of the bacterial culture. Resulting in a 1:10 dilution factor. Then, 1mL of this solution is transferred to the next tube, negative two, resulting in a 1:100 dilution factor. This process continues through the last tube, negative nine, resulting in a final dilution factor of 1:1 billion. After this, 100 microliters of each dilution is plated. The plates are then incubated and the clonal colonies are counted. The dilution plate for a given time point that grows between 30 and 300 colonies is used to calculate the CFUs per milliliter for that time point.
The second common method of measuring bacterial concentration is the optical density. The optical density of a culture can be measured instantly, in relation a media blank, with a spectrophotometer. Typically a wave length of 600 nanometers, also referred to as OD600, is used for these measurements, which increase as cell density increases. While optical density is less precise than CFUs, it is convenient because it can be obtained instantaneously and requires relatively few reagents. Both techniques can be used together to create a standard curve that more accurately approximates the bacterial cell count of a culture. In this video, you will learn how to obtain CFUs and OD600 measurements from timed serial dilutions of E. coli. Then, two growth curves using the CFU and OD600 measurements, respectively, will be plotted before being related by a standard curve.
When working with bacteria, it is important to use the appropriate personal protective equipment such as a lab coat and gloves and to observe proper aseptic technique.
After this, sterilize the work station with 70% ethanol. First, prepare the LB broth and LB solid agar media in separate autoclaveable bottles. After partially closing the caps of the bottles, sterilize the media in an autoclave set to 121 degrees Celsius for 35 minutes. Next, allow the agar media to cool in a water bath set to 50 degrees Celsius for 30 minutes. Once cooled, pour 20 to 25 mL into each Petri dish. After this, allow the plates to set for 24 hours at room temperature.
To prepare the single colony isolates that will later be used to produce a liquid bacterial culture, use previously frozen stock and proper streak plating technique to streak E. coli for isolation on LB agar. Incubate the dish at 37 degree Celsius overnight. After this, cool a flame sterilized inoculation loop on the agar before selecting a single colony from the streaked plate. Inoculate 4 mL of liquid media in a 15 mL test tube. Then, grow the E. coli at 37 degrees Celsius overnight with shaking at 210 rpm.
To set up the 1:1000 volume of bacterial culture that will be used in the growth curve, first obtain an autoclaved 500 mL Erlenmeyer flask. Then, use a 50 mL serological pipette to transfer 100 mL of sterile media to the flask. Next, label nine 15 ml test tubes consecutively as one through nine. These numbers correspond to the dilution factor that will be used to calculate the colony forming unit, or CFU. Then, add 9 mL of 1X PBS to each tube. After this, label the prepared agar plates with the corresponding time points and dilution factors that will be grown. In this example with E. coli, after the starting time point, time points are taken once every hour. Using the previously prepared overnight liquid E. coli culture, inoculate the media in the autoclave 500 mL Erlenmeyer flask with 1:1000 volume of culture. Swirl the media to evenly distribute the bacteria.
After blanking a spectrophotometer, clean the cuvette with a lint-free wipe. Next, dispense 1 mL of the culture into the cuvette and place it into the spectrophotometer to obtain the optical density of the culture at time point zero. Then, grow the E. coli at 37 degrees Celsius with shaking at 210 rpm. At each time point after time point zero, withdraw another 1 mL of bacterial culture from the flask and repeat the optical density measurement. If the optical density reading is greater than 1.0, dilute 100 microliters of bacterial culture with 900 microliters of fresh media and then measure the optical density once more. This value can be multiplied by 10 for the OD 600 measurement.
To obtain the colony forming unit measurement for each time point, withdraw an additional 1 mL of bacterial culture from the flask at each time point. Dispense the bacterial culture into the negative one test tube and vortex to mix. Then, perform the dilution series by first transferring 1 mL from the negative one tube into the negative two tube and vortex to mix. Transfer 1 mL from the negative two tube into the negative three tube and vortex to mix. Continue this serial transfer down all the dilution tubes to the negative nine tube. Dispense 100 microliters of cell suspension onto the correspondingly labeled plate for each dilution. For every dilution, sterilize a cell spreader in ethanol, pass it through a Bunsen burner flame, and cool it by touching the surface of the agar away from the inoculate. Then, use the cell spreader to spread the cell suspension until the surface of the agar plate becomes dry. Incubate the plates upside down at 37 degrees Celsius. Once visible colonies arise, count the number of bacterial colonies on each plate. Record these values and their associated dilution factors for each plate at each time point.
To create an OD 600 growth curve, after ensuring all the data points are entered correctly into a table, select all of the time points and their corresponding data. To generate a colony forming unit growth curve plot, choose the dilution plate where the colony counts fell within the range 30 to 300 bacteria for each time point. Multiply the colony count number by the dilution factor, and then by ten. This is because the 100 microliters spread is considered an additional 1:10 dilution when calculating colony forming units per milliliter. After this, plot the colony forming units versus time on a semi-log scale.
These plots produced with OD 600 and CFU measurements, respectively, can provide valuable information on E. coli growth kinetics. The optical density and colony forming units can be related, so that CFUs per milliliter can be estimated from OD 600 measurements, saving time and materials in future experiments.
To do this, plot the colony forming units against the optical density on a linear scale for OD 600 readings less than or equal to 1. 0. After this, generate a linear regression trend line in Y = MX + B format, where M is the slope and B is the y-intercept. Right click on the data points and select add trend line and linear. Then, check the box to display the equation on the chart and display the R squared value on the chart. The R squared value is the statistical measurement of how closely the data matched the fitted regression line. In this example, the first 6 time points are plotted with OD 600 on the x axis and CFUs per milliliter on the y axis. In future experiments with the same growth conditions, these slope and y-intercept values can be plugged into this equation to estimate CFUs from OD 600 readings. Next, look at the colony forming unit growth curve plot. During the exponential phase, identify two time points with the steepest slope between them. To calculate the doubling time, first calculate the change in time between the selected time points. Then, calculate the change in generations using the equation shown here. Here, lower case b is the number of bacteria at time point three and upper case B is the number of bacteria at time point two. Finally, divide the change in time by the change in generations. In this example, the doubling time is 0. 26 hours or 15 minutes and 19 seconds. Comparing doubling times across different experimental treatment allows us to identify the best growth conditions for a certain bacterial species. Therefore, the treatment with the lowest doubling time will be most optimal of the conditions tested.
