Conjugation: A Method to Transfer Ampicillin Resistance from Donor to Recipient E. coli
English
Share
Overview
Source: Alexander S. Gold1, Tonya M. Colpitts1
1 Department of Microbiology, Boston University School of Medicine, National Emerging Infections Diseases Laboratories, Boston, MA
First discovered by Lederberg and Tatum in 1946, conjugation is a form of horizontal gene transfer between bacteria that relies on direct physical contact between two bacterial cells (1). Unlike other forms of gene transfer, such as transformation or transduction, conjugation is a naturally occurring process in which DNA is secreted from a donor cell to a recipient cell in a unidirectional manner. This directionality and the ability for this process to increase the genetic diversity of bacteria has given conjugation the reputation as a form of bacterial "mating," which is believed to have contributed greatly to the recent rise in antibiotic resistant bacteria (2, 3). By using selective pressures, for example the use of antibiotics, conjugation has been manipulated for use in the laboratory setting, making it a powerful tool for horizontal gene transfer between bacteria, and in some cases from bacteria to yeast, plant, and animal cells (4). Apart from applications in the laboratory, bacterium-eukaryote gene transfer by conjugation is an exciting avenue of DNA transfer with a multitude of possible biotechnical applications and naturally occurring implications (5).
Conjugation is thought to work by a "two-step mechanism" (6). First, before any DNA can be transferred, the donor cell must make direct cell-to-cell contact with the recipient. This process has been characterized best in gram-negative bacteria, the most studied of which is Escherichia coli. Cell-to-cell contact is established by the presence of a complex network of extracellular filaments on the donor known as the sex pilus, a conjugative element encoded for by the transferable gene known as the F (fertility) factor (7, 8). In addition to establishing contact between donor and recipient, several proteins are transported via the sex pilus to the recipient cytoplasm, forming a type IV secretion system (T4SS) conduit between the two cells, a necessary structure for the second step of conjugation, DNA transfer (6). By combining this function of the sex pilus with rolling circle replication of DNA, the donor cell is able to transfer DNA in the form of a transposable element, such as a plasmid or transposon, to the recipient by a "shoot and pump" model (6). In this case, the "shooting" is the transport of the pilot protein, with linked DNA, by the T4SS into the recipient cell, and the "pumping" is the active transport of DNA to the recipient, a process reliant on the T4SS and catalyzed by coupling proteins (6). The machinery used in this this process is composed of an origin of transfer sequence (oriT), which must be provided by the DNA in cis and trans genes, which encode a relaxase, mate pair formation complex, and type IV coupling protein, and can be present in cis or trans (9). This relaxase cleaves the nic site within the oriT sequence and covalently attaches to the 5' end of the transferred strand to produce the relaxosome, a single-stranded DNA-relaxase complex with other auxiliary proteins (9). Once formed, the relaxosome connects to the mating pair formation complex, via the type IV coupling protein, which allows for the transfer of the ssDNA-relaxase complex into recipient cells by the T4SS (10). Once in the cytoplasm of the recipient, the DNA can integrate into the recipient genome or exist separately in the form of a plasmid, either of which allow for the expression of its genes.
In this experiment, the widely used conjugation donor strain E. coli WM3064 was used to transfer the gene encoding for ampicillin resistance to the recipient strain E. coli J53. While both strains of the gram-negative bacteria were resistant to tetracycline, only the donor strain WM3064 had the gene for ampicillin resistance, encoded for in the pWD2-oriT shuttle vector, and was auxotrophic to diaminopimelic acid (DAP) (11-13). This experiment consisted of two main steps, the preparation of donor and recipient strains, followed by the transfer of the ampicillin resistance gene from donor to recipient by conjugation (Figure 1).
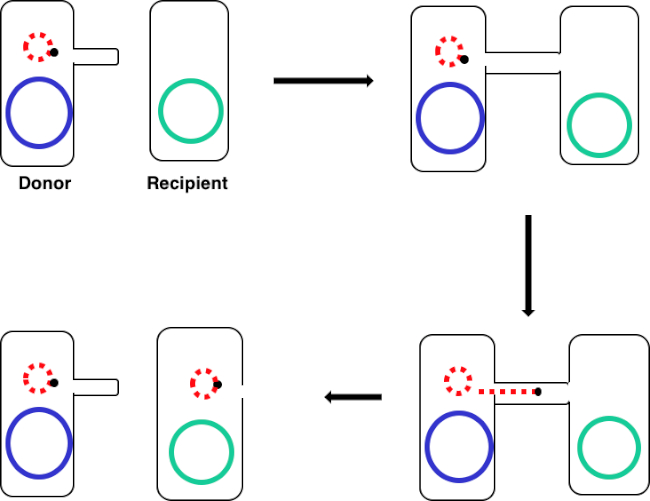
Figure 1: Conjugation schematic. This schematic shows the successful transfer of a plasmid, only one example of a transposable DNA element, from a donor cell to a recipient cell using conjugation. Upon contact with the recipient cell by the donor cell via the sex pilus, the plasmid replicates by rolling circle replication, moves through the multiprotein complex joining the two cells, and forms a new full-length plasmid in the recipient cell.
By incubating a mixture of donor and recipient cells, then successively plating these cells in the presence of tetracycline and DAP, this allowed for the successful transfer of the ampicillin resistance gene. Successively plating cells grown from this mixture in the presence of tetracycline and ampicillin, removed all donor cells due to the lack of DAP and any recipient cells that may not have gained the ampicillin resistance gene, yielding strictly recipient J53 strain bacteria that acquired ampicillin resistance (Figure 2). Once carried out, the successful transfer of the ampicillin resistance gene was confirmed by PCR. Since conjugation was successful, the J53 strain of E. coli contained pWD2-oriT and was resistant to ampicillin, and the gene encoding for this resistance is detectable by PCR. However, if unsuccessful there would have been no detection of the ampicillin resistance gene and ampicillin would still function as an effective antibiotic against the J53 strain.
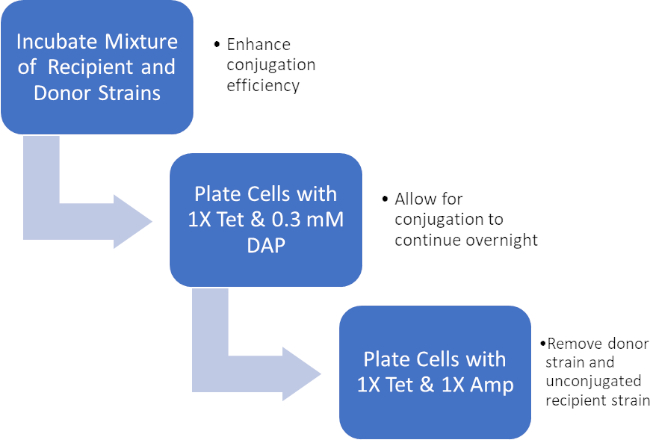
Figure 2: Protocol schematic. This schematic shows an overview of the presented protocol.
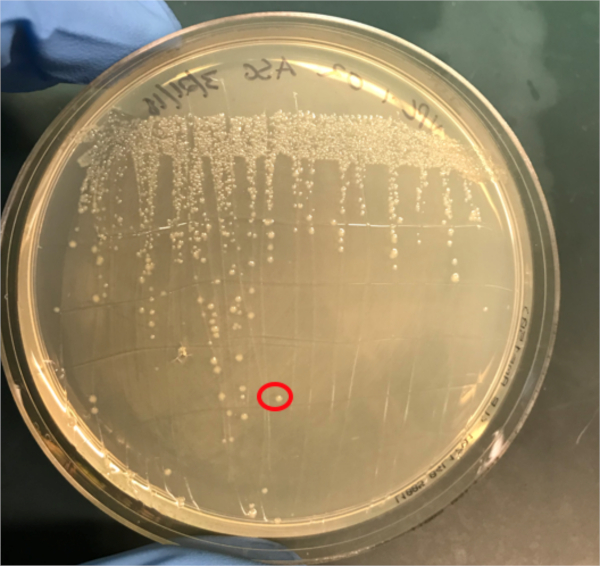
Figure 3A: The confirmation of successful conjugation by PCR. A) Freezer stocks of the conjugated and negative control samples were streaked out on agar plates and a colony was selected (red) for DNA isolation.
Procedure
1. Set-Up
- Autoclave approximately 1L of Luria-Bertani medium (LB). This sterile LB will be used to make approximately 5 mL of LB containing 0.3 mM diaminopimelic acid (DAP).
- Collect the following plates: LB agar plates with 1X Tet and 0.3 mM DAP, LB agar plates with 1X Tet only, and LB agar plates with 1X Amp/Tet only.
- Ensure that some glycerol and a box of pre-sterilized plastic pipette tips are close at hand.
- Prior to starting any work involving microbes, sterilize the workspace with 70% ethanol. Always wear the necessary personal protective equipment, including a lab coat and gloves.
- Once finished, sterilize all surfaces and gloves with 70% ethanol and wash hands.
2. Donor and Recipient Strain Preparation
- Prepare 5 mL bacterial cultures of the donor and recipient strains and grow them overnight at 37 °C with aeration and shaking at 220 rpm. The donor strain should be grown in LB with 0.3 mM DAP.
- Spin down 1 mL of both cultures (~3000 rpm for 5 minutes) and wash the cells with PBS.
- Resuspend the cells in 500 µL of phosphate buffered saline (PBS).
3. Conjugation
- Combine 50 µL of recipient cells with 50 µL of donor cells in a microcentrifuge tube and mix by pipetting.
- Incubate the cell mixture for 1 hour at 37 °C. This will enhance the efficiency of conjugation before and during the next plating step.
- Pipet 100 µL of the cell mixture onto an agar plate with 1X Tet and 0.3 mM DAP. Do not spread on the plate.
- Pipet 100 µL of recipient cell culture onto an agar plate with 1X Tet and 0.3 mM DAP. Do not spread on the plate.
- Incubate both plates overnight at 37 °C.
- Scrape up the conjugation cell mixture and the recipient cell culture with a sterile cell scraper. Transfer the cells to sterile microcentrifuge tube and resuspend the cells in 1 mL of PBS.
- Vortex the cells and gently spin them down (~3000 rpm for 5 minutes).
- Resuspend the cells in 1 mL of PBS.
- Plate the conjugation reaction cell mixture on an LB agar plate with 1X Amp/Tet only. On this plate, only the recipient bacteria that successfully gained the Amp resistance gene via conjugation are expected to grow.
- Plate the recipient cells onto an LB agar plate with 1X Tet only. On this plate, only unconjugated recipient bacteria, which do not have the Amp resistance gene, are expected to grow.
- Incubate the plates overnight at 37 °C.
- Pick single colonies from both plates and use them to grow overnight cultures in 5 mL of media (37 °C with aeration at 220 rpm).
4. DNA Isolation
- Isolate DNA from the previously prepared cultures using 4.5 mL of the total culture volume by DNA miniprep.
- To do this, elute the DNA using 35 µL of nuclease-free water.
- Pure DNA will generate an absorbance ratio (A260/280) of approximately 1.8.
- Use the remaining 0.5 mL of each culture to prepare glycerol stocks by making a 1:1 mixture of bacterial culture and 100% glycerol.
- Store the freezer stocks at -80 °C.
5. Confirmation of Plasmid Transfer Conjugation by PCR
- Prepare two PCR master mixes, each with a different set of forward and reverse primers, one targeting a 500 base-pair segment within the ampicillin resistance gene and the other targeting a segment within a housekeeping gene.
- Housekeeping gene primers were designed to amplify a segment of DNA within the bacterial gene encoding for DNA gyrase B (14).
- The following reagent volumes were used to prepare 90 μL of each master mix:
7.5 μL of 10 μM forward primer
7.5 μL of 10 μM reverse primer
75 μL of 2X PCR Master Mix
- Prepare the following six PCR reactions using 15 μL master mix, 10 ng of template DNA, and nuclease-free water up to a final volume of 25 μL.
Conjugation reaction DNA and ampicillin primers
Conjugation reaction DNA and housekeeping primers
Negative control DNA and ampicillin primers
Negative control DNA and housekeeping primers
No DNA and ampicillin primers
No DNA and housekeeping primers - Transfer these reactions to a PCR machine with the block preheated to 98 C and begin thermocycling under the following conditions:
98 °C for 30 seconds
25-35 cycles of 98 °C for 5-10 seconds, 45-72 °C for 10 to 30 seconds, and 72 °C for 15-30 seconds per kb
72 °C for 5-10 minutes
Hold at 4 °C - Load all six PCR reactions onto a 1% agarose gel and run at ~150V for about 20 minutes.
- Visualize the PC product using a UV illuminator.
Bacterial cells, such as E. coli, are able to transfer genetic information from cell-to-cell. Conjugation differs from other mechanisms of DNA transfer, such as transduction or transformation, in that it requires physical contact between the cells.
To proceed, conjugation requires a donor cell that expresses the fertility, or F, factor and a recipient cell without it, an F minus cell. The process requires two steps. The first is the establishment of direct cell-to-cell contact. To do this, the donor cell generates an extracellular filamentous structure called a sex pilus. It is named this since conjugation is a form of mating for asexually reproducing bacteria, but it should be noted that it is not true sexual reproduction as no gametes are exchanged and no offspring are formed.
The second step is delivery of DNA to the recipient cell. After the sex pilus establishes contact between two cells, a conduit called the Type IV secretion system is built allowing for the transfer of DNA. The donor cell then begins to replicate the extrachromosomal DNA that will be transferred selected based on the presence of a genetic element known as the OriT or origin of transfer. One end of the newly replicated DNA is threaded into the conduit through DNA protein binding. As the DNA is further replicated, it is pumped through the channel, facilitated by a complex of proteins encoded by genes located close to the OriT. Once the DNA is fully transferred, it will either form an extra chromosomal plasmid, or it may integrate into the chromosome of the recipient cell. Whichever the endpoint of the transferred DNA, the genes it encodes will then be expressed. This gene expression can be used to confirm successful conjugation.
For example, consider a scenario where the donor strain expresses ampicillin resistance and passes this on in the conjugated DNA to the recipient bacterium, but the recipient strain also has a tetracycline resistance gene not present in the donor. In this event, when the cells are plated on LB media containing both tetracycline and ampicillin, colonies should grow only from successfully conjugated bacteria, which will be expressing both resistance phenotypes. To further confirm successful conjugation, plasmid DNA from these colonies can be harvested and then a section of DNA specific to the transferred plasmid can be amplified using polymerase chain reaction, or PCR. When the PCR product is run on an electrophoresis gel alongside a ladder of standard sizes, a PCR fragment of a known size should be visible on the gel, further confirming successful conjugation. In this experiment, a plasmid will be used to transfer the ampicillin resistance gene via conjugation from a donor strain to a tetracycline-resistant recipient strain. After this, to confirm conjugation, the conjugation mixture will be incubated on a plate containing both antibiotics leaving only the transformed bacteria. Finally, successful conjugation will be further confirmed with PCR.
Before starting the procedure, put on the appropriate personal protective equipment, including a lab coat and gloves. Next, sterilize the workspace using 70% ethanol to wipe down the surface.
In this procedure, the ampicillin resistance gene will be transferred from the WM3064 strain of E. coli to the J53 strain of E. coli via conjugation. The donor strain WM3064 is resistant to tetracycline and ampicillin and it requires diaminopimelic acid, or DAP, to grow. The recipient strain J53 is only resistant to tetracycline and it does not require DAP to grow. This means that successfully conjugated cells should be resistant to tetracycline and ampicillin and can grow without DAP.
Prepare the donor strain culture by inoculating five milliliters of LB containing 0.3 millimoles of DAP with a scrap of the frozen donor strain glycerol stock. Then, prepare the recipient strain by inoculating five milliliters of LB broth without DAP with a scrap of the frozen recipient strain glycerol stock. Grow these cultures overnight at 37 degrees Celsius with aeration and shaking at 220 RPM in a shaking incubator. Once the cultures have grown to an OD 600 of two, remove one milliliter of culture from each and place this into two new separate 1.5 milliliter microcentrifuge tubes. Then, centrifuge these aliquots at 3000 RPM for five minutes to pellet the bacterial cells. Discard the supernatant and wash each pellet with 250 microliters of 1X PBS. Centrifuge the samples again and, after discarding the supernatant, resuspend each pellet in 500 microliters of PBS.
To begin the conjugation procedure, first combine 50 microliters of recipient cells with 50 microliters of donor cells in a 1.5 milliliter microcentrifuge tube and mix by pipetting up and down gently. Next, pipette 100 microliters of the recipient cell culture onto another 1X tetracycline plate containing DAP. Next, prepare your negative control by pipetting 100 microliters of the recipient cell culture only onto a non-selective agar plate containing DAP. Then, incubate the conjugation and negative control plates overnight at 37 degrees Celsius.
The next day, take a sterile cell scraper and harvest cells from the conjugation plate by collecting colonies. Then, transfer the colonies to a sterile 1.5 milliliter microcentrifuge tube containing one milliliter of 1X PBS. Repeat this process to collect the recipient cells from the other plate.
After this, vortex the samples to mix. After mixing, transfer the tubes to a centrifuge to gently pellet the cells. Discard the supernatant, then wash the cell pellets in one milliliter of PBS and vortex the tubes to resuspend the cells. Pellet the cells again by centrifuging. Discard the supernatant again and resuspend both cell pellets in one milliliter of PBS. Now, using a sterile pipette tip, plate 100 microliters of the conjugation reaction cell mixture onto an LB agar plate without DAP containing 1X tetracycline and 1X ampicillin. Repeat the plating method using 100 microliters of a ten-fold dilution of the same cell mixture in PBS onto another LB agar plate without DAP containing 1X tetracycline and 1X ampicillin.
Finally, pipette 100 microliters of the negative control cell mixture onto a single LB agar plate with 1X tetracycline only. After overnight incubation at 37 degrees Celsius, the colonies should be visible. Using a sterile pipette tip, pick a single colony from the conjugation reaction plate and add it to a tube containing five milliliters of selective LB media containing both antibiotics. Then, repeat the colony isolation by selecting a single colony from the recipient cell plate. Grow these cultures overnight at 37 degrees Celsius with aeration at 220 RPM.
The next day, wipe down the bench top with 70% ethanol and remove the plates from the incubator. Use a DNA mini prep kit to isolate DNA from 4. 5 milliliters of each culture according to the manufacturer's instructions. After completing the DNA mini prep, elute the DNA using 35 microliters of nuclease-free water. Finally, use the remaining 0. 5 milliliters of each culture to prepare one milliliter glycerol stocks by adding 0.5 milliliters of 100% glycerol for a one-to-one dilution. Place these aliquots at minus 80 degrees Celsius for storage until needed.
To confirm successful conjugation by PCR, first prepare a PCR master mix by adding 75 microliters of 2X PCR master mix to a microcentrifuge tube. Then, add 7.5 microliters each of a 10 micromolar forward primer and a 10 micromolar reverse primer designed to amplify the ampicillin resistance gene from the plasmid. Next, prepare a second PCR master mix by adding 75 microliters of 2X PCR master mix to a microcentrifuge tube and then adding 7.5 microliters each of a 10 micromolar forward primer and 10 micromolar reverse primer designed to amplify a housekeeping gene, in this case DNA gyrase B.
Now, add 15 microliters of the first master mix to a PCR tube and then add 10 nanograms, approximately two microliters of the template experimental DNA to the same tube. Bring the reaction up to a final volume of 25 microliters with nuclease-free water. Repeat these steps to produce the remaining five reactions, so that the tubes contain the components shown here. Now, transfer these reactions to a thermocycler with the block pre-heated to 98 degrees Celsius and then initiate the program. After completion of the PCR, remove the tubes from the machine. Then, load two microliters of each reaction mixed with two microliters of loading dye and four microliters of a molecular weight marker into consecutive wells of a 1% agarose gel. Set the gel to run at 150 volts for 20 minutes. Finally, visualize the gel using a UV illuminator.
In this experiment, the successful transfer of the ampicillin resistance gene via conjugation was confirmed via PCR. Here, a roughly 500 base pair sized band should be observed in the well containing the conjugated DNA and ampicillin primers, well two in this example. A housekeeping gene, DNA gyrase B, was loaded into wells three and five with conjugated DNA and recipient cell DNA, respectively. Bands observed in these wells act as a positive control to ensure the DNA template was present and that PCR was successful. Bands should not be observed in the well containing the reaction for recipient cell DNA and the ampicillin primer pair, well four in this example, because the recipient cells are not ampicillin-resistant. Additionally, no bands should be observed in the reactions lacking template DNA, wells six and seven here. If these conditions are met, this will confirm the successful transfer of the ampicillin resistance gene, conferring ampicillin resistance from the WM3064 strain of E. coli to the J53 strain of E. coli.
Results
If conjugation was successful, a 500 base-pair sized band PCR product will be observed in the well in which PCR reaction 1 was loaded (Well #2 in Figure 3B), while no bands will be observed in the well in which PCR reaction 3 was loaded (Well #4 in Figure 3B). The presence of this band confirms the successful transfer of the ampicillin resistance gene, thereby conferring ampicillin resistance to the J53 strain of E. coli.

Figure 3B: The confirmation of successful conjugation by PCR. B) PCR analysis was done using DNA isolated from the select colony. The contents of each well are as follows: 1) DNA ladder, 2) Conjugation DNA and ampicillin primers, 3) Conjugation DNA and housekeeping primers, 4) Negative control DNA and ampicillin primers, 5) Negative control DNA and housekeeping primers, 6) No DNA and ampicillin primers, and 7) No DNA and negative control primers. The presence of a ~ 500 base-pair band PCR product from PCR reaction 1 (well 2), and the lack of this product from PCR reaction 3 (well 4), confirms successful conjugation.
Applications and Summary
Conjugation is a naturally occurring process of horizontal gene transfer that relies on the direct cell-to-cell contact of a donor cell and a recipient cell. This process is shared among all kinds of bacteria and has been instrumental in bacterial evolution, most notably antibiotic resistance. In the lab, conjugation can be used as an effective method of gene transfer that is much less disruptive when compared to other techniques. Outside of the laboratory, the ability to transfer DNA from bacteria to eukaryotes via conjugation offers an exciting new avenue of gene therapy and understanding the implications of these naturally occurring gene transfers, for example the relationship between bacterial infection and cancer, is a rapidly emerging area of research.
References
- Lederberg J, Tatum, E.L. Gene recombination in Escherichia coli Nature. 1946;158:558.
- Holmes R.K. J, M.G. Genetics: Exchange of Genetic Information. 4th Edition ed. Baron S, editor. Galveston, TX: University of Texas Medical Branch at Galveston; 1996.
- Cruz F, Davies, J. Horizontal gene transfer and the origin of species: lessons from bacteria. Trends in Microbiology. 2000;8:128-33.
- Llosa M, Cruz, F. Bacterial conjugation: a potential tool for genomic engineering. Ressearch in Microbiology. 2005;156:1-6.
- Lacroix B, Citovsky, V. Transfer of DNA from Bacteria to Eukaryotes. mBio. 2016;7(4):1-9.
- Llosa M, et al. Bacterial conjugation: a two-step mechanism for
- DNA transport. Molecular Microbiology. 2002;45:1-8.
- Grohmann E, Muth, G., Espinosa, M. Conjugative Plasmid Transfer in Gram-Positive Bacteria. Microbiology and Molecular Biology Reviews. 2003;67:277-301.
- Firth N, Ippen-Ihler, K, Skurray, RA. Structure and function of the F factor and mechanism of conjugation. Escherichia coli and salmonella: cellular and molecular biology. 1996;2:2377-401.
- Smillie C, Garcillan-Barcia MP, Francia MV, Rocha EPC, De La Cruz F. Mobility of Plasmids. Microbiology and Molecular Biology Reviews. 2010;74(3):434-52.
- Cascales E. Definition of a Bacterial Type IV Secretion Pathway for a DNA Substrate. 2004;304(5674):1170-3.
- Wang P, Yu Z, Li B, Cai X, Zeng Z, Chen X, et al. Development of an efficient conjugation-based genetic manipulation system for Pseudoalteromonas. Microbial Cell Factories. 2015;14(1):11.
- Yi H, Cho YJ, Yong D, Chun J. Genome Sequence of Escherichia coli J53, a Reference Strain for Genetic Studies. Journal of Bacteriology. 2012;194(14):3742-3.
- Baumann RLB, E. H.; Wiseman, J. S.; Vaal, M.; Nichols, J. S. Inhibition of Escherichia coli Growth and Diaminopimelic Acid Epimerase by 3-Chlorodiaminopimelic Acid. Antimicrobial Agents and Chemotherapy 1988;32:1119-23.
- Rocha D, Santos, CS, Pacheco LG. Bacterial reference genes for gene expression studies by RT-qPCR: survey and analysis. Antonie Van Leeuwenhoek. 2015;108:685-93.
Transcript
Bacterial cells, such as E. coli, are able to transfer genetic information from cell-to-cell. Conjugation differs from other mechanisms of DNA transfer, such as transduction or transformation, in that it requires physical contact between the cells.
To proceed, conjugation requires a donor cell that expresses the fertility, or F, factor and a recipient cell without it, an F minus cell. The process requires two steps. The first is the establishment of direct cell-to-cell contact. To do this, the donor cell generates an extracellular filamentous structure called a sex pilus. It is named this since conjugation is a form of mating for asexually reproducing bacteria, but it should be noted that it is not true sexual reproduction as no gametes are exchanged and no offspring are formed.
The second step is delivery of DNA to the recipient cell. After the sex pilus establishes contact between two cells, a conduit called the Type IV secretion system is built allowing for the transfer of DNA. The donor cell then begins to replicate the extrachromosomal DNA that will be transferred selected based on the presence of a genetic element known as the OriT or origin of transfer. One end of the newly replicated DNA is threaded into the conduit through DNA protein binding. As the DNA is further replicated, it is pumped through the channel, facilitated by a complex of proteins encoded by genes located close to the OriT. Once the DNA is fully transferred, it will either form an extra chromosomal plasmid, or it may integrate into the chromosome of the recipient cell. Whichever the endpoint of the transferred DNA, the genes it encodes will then be expressed. This gene expression can be used to confirm successful conjugation.
For example, consider a scenario where the donor strain expresses ampicillin resistance and passes this on in the conjugated DNA to the recipient bacterium, but the recipient strain also has a tetracycline resistance gene not present in the donor. In this event, when the cells are plated on LB media containing both tetracycline and ampicillin, colonies should grow only from successfully conjugated bacteria, which will be expressing both resistance phenotypes. To further confirm successful conjugation, plasmid DNA from these colonies can be harvested and then a section of DNA specific to the transferred plasmid can be amplified using polymerase chain reaction, or PCR. When the PCR product is run on an electrophoresis gel alongside a ladder of standard sizes, a PCR fragment of a known size should be visible on the gel, further confirming successful conjugation. In this experiment, a plasmid will be used to transfer the ampicillin resistance gene via conjugation from a donor strain to a tetracycline-resistant recipient strain. After this, to confirm conjugation, the conjugation mixture will be incubated on a plate containing both antibiotics leaving only the transformed bacteria. Finally, successful conjugation will be further confirmed with PCR.
Before starting the procedure, put on the appropriate personal protective equipment, including a lab coat and gloves. Next, sterilize the workspace using 70% ethanol to wipe down the surface.
In this procedure, the ampicillin resistance gene will be transferred from the WM3064 strain of E. coli to the J53 strain of E. coli via conjugation. The donor strain WM3064 is resistant to tetracycline and ampicillin and it requires diaminopimelic acid, or DAP, to grow. The recipient strain J53 is only resistant to tetracycline and it does not require DAP to grow. This means that successfully conjugated cells should be resistant to tetracycline and ampicillin and can grow without DAP.
Prepare the donor strain culture by inoculating five milliliters of LB containing 0.3 millimoles of DAP with a scrap of the frozen donor strain glycerol stock. Then, prepare the recipient strain by inoculating five milliliters of LB broth without DAP with a scrap of the frozen recipient strain glycerol stock. Grow these cultures overnight at 37 degrees Celsius with aeration and shaking at 220 RPM in a shaking incubator. Once the cultures have grown to an OD 600 of two, remove one milliliter of culture from each and place this into two new separate 1.5 milliliter microcentrifuge tubes. Then, centrifuge these aliquots at 3000 RPM for five minutes to pellet the bacterial cells. Discard the supernatant and wash each pellet with 250 microliters of 1X PBS. Centrifuge the samples again and, after discarding the supernatant, resuspend each pellet in 500 microliters of PBS.
To begin the conjugation procedure, first combine 50 microliters of recipient cells with 50 microliters of donor cells in a 1.5 milliliter microcentrifuge tube and mix by pipetting up and down gently. Next, pipette 100 microliters of the recipient cell culture onto another 1X tetracycline plate containing DAP. Next, prepare your negative control by pipetting 100 microliters of the recipient cell culture only onto a non-selective agar plate containing DAP. Then, incubate the conjugation and negative control plates overnight at 37 degrees Celsius.
The next day, take a sterile cell scraper and harvest cells from the conjugation plate by collecting colonies. Then, transfer the colonies to a sterile 1.5 milliliter microcentrifuge tube containing one milliliter of 1X PBS. Repeat this process to collect the recipient cells from the other plate.
After this, vortex the samples to mix. After mixing, transfer the tubes to a centrifuge to gently pellet the cells. Discard the supernatant, then wash the cell pellets in one milliliter of PBS and vortex the tubes to resuspend the cells. Pellet the cells again by centrifuging. Discard the supernatant again and resuspend both cell pellets in one milliliter of PBS. Now, using a sterile pipette tip, plate 100 microliters of the conjugation reaction cell mixture onto an LB agar plate without DAP containing 1X tetracycline and 1X ampicillin. Repeat the plating method using 100 microliters of a ten-fold dilution of the same cell mixture in PBS onto another LB agar plate without DAP containing 1X tetracycline and 1X ampicillin.
Finally, pipette 100 microliters of the negative control cell mixture onto a single LB agar plate with 1X tetracycline only. After overnight incubation at 37 degrees Celsius, the colonies should be visible. Using a sterile pipette tip, pick a single colony from the conjugation reaction plate and add it to a tube containing five milliliters of selective LB media containing both antibiotics. Then, repeat the colony isolation by selecting a single colony from the recipient cell plate. Grow these cultures overnight at 37 degrees Celsius with aeration at 220 RPM.
The next day, wipe down the bench top with 70% ethanol and remove the plates from the incubator. Use a DNA mini prep kit to isolate DNA from 4. 5 milliliters of each culture according to the manufacturer’s instructions. After completing the DNA mini prep, elute the DNA using 35 microliters of nuclease-free water. Finally, use the remaining 0. 5 milliliters of each culture to prepare one milliliter glycerol stocks by adding 0.5 milliliters of 100% glycerol for a one-to-one dilution. Place these aliquots at minus 80 degrees Celsius for storage until needed.
To confirm successful conjugation by PCR, first prepare a PCR master mix by adding 75 microliters of 2X PCR master mix to a microcentrifuge tube. Then, add 7.5 microliters each of a 10 micromolar forward primer and a 10 micromolar reverse primer designed to amplify the ampicillin resistance gene from the plasmid. Next, prepare a second PCR master mix by adding 75 microliters of 2X PCR master mix to a microcentrifuge tube and then adding 7.5 microliters each of a 10 micromolar forward primer and 10 micromolar reverse primer designed to amplify a housekeeping gene, in this case DNA gyrase B.
Now, add 15 microliters of the first master mix to a PCR tube and then add 10 nanograms, approximately two microliters of the template experimental DNA to the same tube. Bring the reaction up to a final volume of 25 microliters with nuclease-free water. Repeat these steps to produce the remaining five reactions, so that the tubes contain the components shown here. Now, transfer these reactions to a thermocycler with the block pre-heated to 98 degrees Celsius and then initiate the program. After completion of the PCR, remove the tubes from the machine. Then, load two microliters of each reaction mixed with two microliters of loading dye and four microliters of a molecular weight marker into consecutive wells of a 1% agarose gel. Set the gel to run at 150 volts for 20 minutes. Finally, visualize the gel using a UV illuminator.
In this experiment, the successful transfer of the ampicillin resistance gene via conjugation was confirmed via PCR. Here, a roughly 500 base pair sized band should be observed in the well containing the conjugated DNA and ampicillin primers, well two in this example. A housekeeping gene, DNA gyrase B, was loaded into wells three and five with conjugated DNA and recipient cell DNA, respectively. Bands observed in these wells act as a positive control to ensure the DNA template was present and that PCR was successful. Bands should not be observed in the well containing the reaction for recipient cell DNA and the ampicillin primer pair, well four in this example, because the recipient cells are not ampicillin-resistant. Additionally, no bands should be observed in the reactions lacking template DNA, wells six and seven here. If these conditions are met, this will confirm the successful transfer of the ampicillin resistance gene, conferring ampicillin resistance from the WM3064 strain of E. coli to the J53 strain of E. coli.
