The Submerged Printing of Cells onto a Modified Surface Using a Continuous Flow Microspotter
Summary
This 3D microfluidic printing technology prints arrays of cells onto submerged surfaces. We describe how arrays of cells are delivered microfluidically in 3D flow cells onto submerged surfaces. By printing onto submerged surfaces, cell microarrays were produced that allow for drug screening and cytotoxicity assessment in a multitude of areas.
Abstract
The printing of cells for microarray applications possesses significant challenges including the problem of maintaining physiologically relevant cell phenotype after printing, poor organization and distribution of desired cells, and the inability to deliver drugs and/or nutrients to targeted areas in the array. Our 3D microfluidic printing technology is uniquely capable of sealing and printing arrays of cells onto submerged surfaces in an automated and multiplexed manner. The design of the microfluidic cell array (MFCA) 3D fluidics enables the printhead tip to be lowered into a liquid-filled well or dish and compressed against a surface to form a seal. The soft silicone tip of the printhead behaves like a gasket and is able to form a reversible seal by applying pressure or backing away. Other cells printing technologies such as pin or ink-jet printers are unable to print in submerged applications. Submerged surface printing is essential to maintain phenotypes of cells and to monitor these cells on a surface without disturbing the material surface characteristics. By printing onto submerged surfaces, cell microarrays are produced that allow for drug screening and cytotoxicity assessment in a multitude of areas including cancer, diabetes, inflammation, infections, and cardiovascular disease.
Introduction
Recent advances in the pharmaceutical industry have led to increased interest in using cellular microarrays in the drug discovery process for drug screening and cytotoxicological analysis1,2,3. The development of in vitro high-throughput assays and screening methods using cell microarrays would facilitate the rapid and cost-effective development of drug candidates as well as advance the fundamental understanding of the cell1,4. The traditional approach to screening with cells uses conventional well-plate platforms; however this approach is limited due to the high cost, limited throughput, and limited ability for quantitative information on cell function1,5. Due to these limitations, research in cellular microarray technologies is burgeoning for molecular biological characterization, tissue engineering, and drug screening1,6. The advantages of cellular microarrays include smaller sample use, minimal effects of cellular phenotype heterogeneity masking information, and most importantly the ability to automate assays for more high-throughput applications1,7,8.
The pharmaceutical industry currently utilizes high-throughput cell-based screening assays with 2D cell monolayer cultures for drug screening in microtiter well plates9. Multiplexing cells in wells of microtiter plates offers the potential for higher throughput with unique experimentation options. Further, the current technologies for cellular microarrays allow the cells to dry which could dramatically alter the phenotype of the cells from in vivo10,11. In order to overcome these problems, the MFCA was engineered and is shown in Figure 1. The design of the MFCA 3D fluidics enables the printhead tip in Figure 1 to be lowered into a bath and compressed against a surface to form a seal. The soft silicone tip of the printhead behaves like a gasket and forms a reversible seal. The MFCA technology is uniquely suited to interface with submerged surfaces, which is required for both cell cultures and tissue slice systems, and is difficult or impossible with most other approaches. Pins or ink-jet printing will not work, and 2D microfluidic devices are not suited for deposition or interfacing with large arrays of discrete spots. Further, by miniaturizing and localizing the experiment – the cellular microarray – the MFCA overcomes the major problems associated with high-throughput cell-based screening assays.
The CFM uses 3D channel networks to cycle small volume fluid samples over microscopic spot locations on a surface12,13. By printing with flow, biomolecules, cells, and other reagents are maintained in a liquid environment throughout the printing process, enabling the printing of sensitive biomolecules and cells without exposure to air, which hinders the current cell printing techniques. It is also possible to print directly from crude material such as hybridoma or supernatants provided there is a capture mechanism on the array surface. The objective of this manuscript is to explain in detail the submerged printing of two cell types onto a surface.
Protocol
1. Cell Culture Preparation
- Store NIH/3T3 cell stocks in liquid nitrogen until ready for use.
- Prepare complete media for NIH/3T3 cells using Dulbecco’s Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum, 10 mM HEPES buffer, 50 units/ml penicillin, and 50 μg/ml streptomycin.
- Thaw cells for 2-3 min in a shaking water bath at 37 °C.
- Resuspend cells in 5 ml of complete media and centrifuge at 1,500 x g for 3 min.
- Remove the cell supernatant without disturbing the cell pellet.
- Resuspend the cells in 5 ml of media and count cells using a hemocytometer.
- Remove 10 μl of the cell suspension and place in a 0.5 ml microcentrifuge tube.
- Add 10 μl of Trypan Blue to the cell suspension and stir with the tip of the pipette.
- Pipette 10 μl of the stained cells into a hemocytometer chamber.
- Count cells in hemocytometer at 10x magnification.
- Count the live cells (small, white balls) for 4 of the 9 large squares. Only count the live cells that do not appear dark blue from the Trypan Blue stain.
- Calculate the average number of cells per large square, resulting in the number of cells x 104 cells/ml in the original cells suspension.
- Seed cells at a density of 1 x105 cells/ml with a total of 5 ml in each T25 flask.
- Culture cells to between 70% and 80% confluency before beginning experiments. Refresh media every 2-3 days or as needed.
2. Printing Surface Preparation
- Mark a rectangle with marker on the bottom of a tissue-culture-treated polystyrene (TCTPS) 12 well plate the size of the print head (7 mm x 19 mm).
- Pipette 50 μl of fetal bovine serum into the rectangular area and spread it over the entire region with the tip.
- Leave the Petri dish in a sterile biosafety hood O/N to allow the serum spot to dry completely. These plates should be prepared no more than 2 days prior to use.
3. Submerged Printing
- Rinse the print head out with distilled water.
- Fill a 60 mm Petri dish with distilled water and dock the print head onto the surface.
- Flow distilled water through each of the lines for 2 min at 150 μl/min using a pneumatic pump.
- Discard the water from the Petri dish and fill it with clean water.
- Move the print head up and down in the water 3 times.
- Dock the print head center over the serum-coated spot in the pre-prepared 12 well plate.
- Prime the print head with pre-warmed complete media, 300 μl per channel.
- Print suspended cells at a concentration of 50,000 cells/ml onto the surface and 60 μl/min, 100 μl per channel.
- Place the print head, left docked against the surface, and the manifold in a cell culture incubator set to 5% CO2 and 37 °C for 2 hr.
- After the 2 hr incubation, remove the print head and place the lid on the culture dish. The time at which the print head removed is considered the 0 hr time point.
4. Standard Cell Culture
- Add 1 ml of pre-warmed media to one of the serum-spotted TCTPS 12-well plate prepared in step 2.
- Take 1 ml of the remaining cell suspension from step 2 and pipette into a single well of a 12-well TCTPS plate. This will produce a culture containing 50,000 cells in the dish.
- Place the cell culture plate in an incubator at 37 °C for 2 hr.
- After the 2 hr incubation begin timing; for consistency between printed and seeding samples, this is considered the 0 hr time point.
5. Culture Visualization
- After 0, 2, 24, and 48 hr take cells from each of the two methods described above and image using a standard inverted microscope.
- Stain the cells using propidium iodide, a red fluorescent stain that is only incorporated into the nucleus of dead cells.
- Gently rinse the cells 3 times with 1 ml of phosphate buffered saline with calcium and magnesium (PBS++) to remove any debris.
- Add 1 ml of pre-warmed PBS++ to each culture vessel.
- Add 1 μl of propidium iodide stock solution (1 mg/ml) to each culture vessel.
- Incubate the cells stained with propidium iodide at 37 °C for 10 min.
- Image the cells using an inverted microscope at 10x and 40x magnification.
- Stain the cells using propidium iodide, a red fluorescent stain that is only incorporated into the nucleus of dead cells.
- Discard the cells in an appropriate biohazard container.
Representative Results
The fibroblast cell line NIH/3T3 cells were printed or seeded onto a submerged surface. The cells were grown to a density of 5 X 104 cells per ml. Cells were seeded using traditional cell culture techniques. Cells were printed using a large-format, twelve-flow cell printhead where the channels are larger (~500 µm) than the CFM used for proteins and other biomolecules. The cells were printed or seeded onto a serum coated surface. The printing process is shown in Figure 1.
After printing and seeding, the cells were visualized to assess density and morphology at relevant time points (0, 2, 24, and 48 hr). The cell density and morphology of the cells at these time points was assessed using 10X and 40X magnifications microscopically. The images show that the cells possess almost identical phenotypes at each time point and at each magnification Figure 2. Based on these figures, the effect of printing was determined to be minimal.
The cell viability was assessed using a PI stain. As shown in Figure 3, the cell viability for the printed cell was not significantly different than the seeded cells for each time point. The major difference between the two methods of attaching the cells to the surface is the density of the cells printed. Cell density decreases at the 2 hr time point by over 50% in both seeded and printed cells. Further study is warranted to determine the cause of this decrease; however, overall printed density ratios when compared to seeded remains significantly higher. The printed cell density in the flowcell was approximately ten times as dense at each time point as the seeded cells considering cells were printed at the same density as seeding, but in a smaller area (0.6 cm2 area for a flow cell, 3.8 cm2 for a well of a 24-well plate). At the final time point the cells are at the same density which is likely due to cell motility and resource consumption limitations.
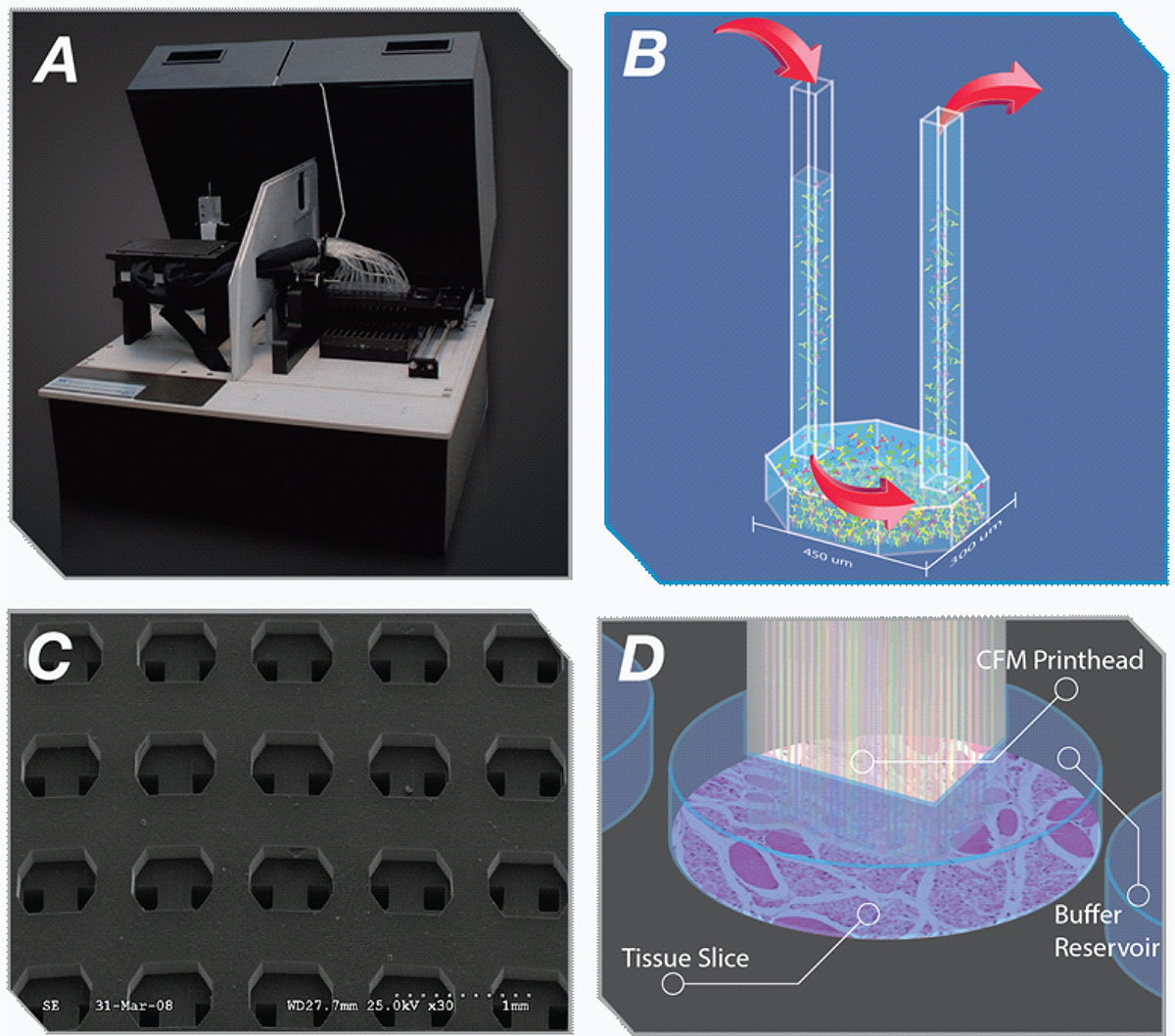
Figure 1. (A) Picture of the CFM. (B) Schematic of a flow cell. (C) SEM Image of the printhead showing individual flowcells. (D) Schematic of submerged printing where printhead docks to tissue culture compatible surface. Please click here to view a larger version of this figure.
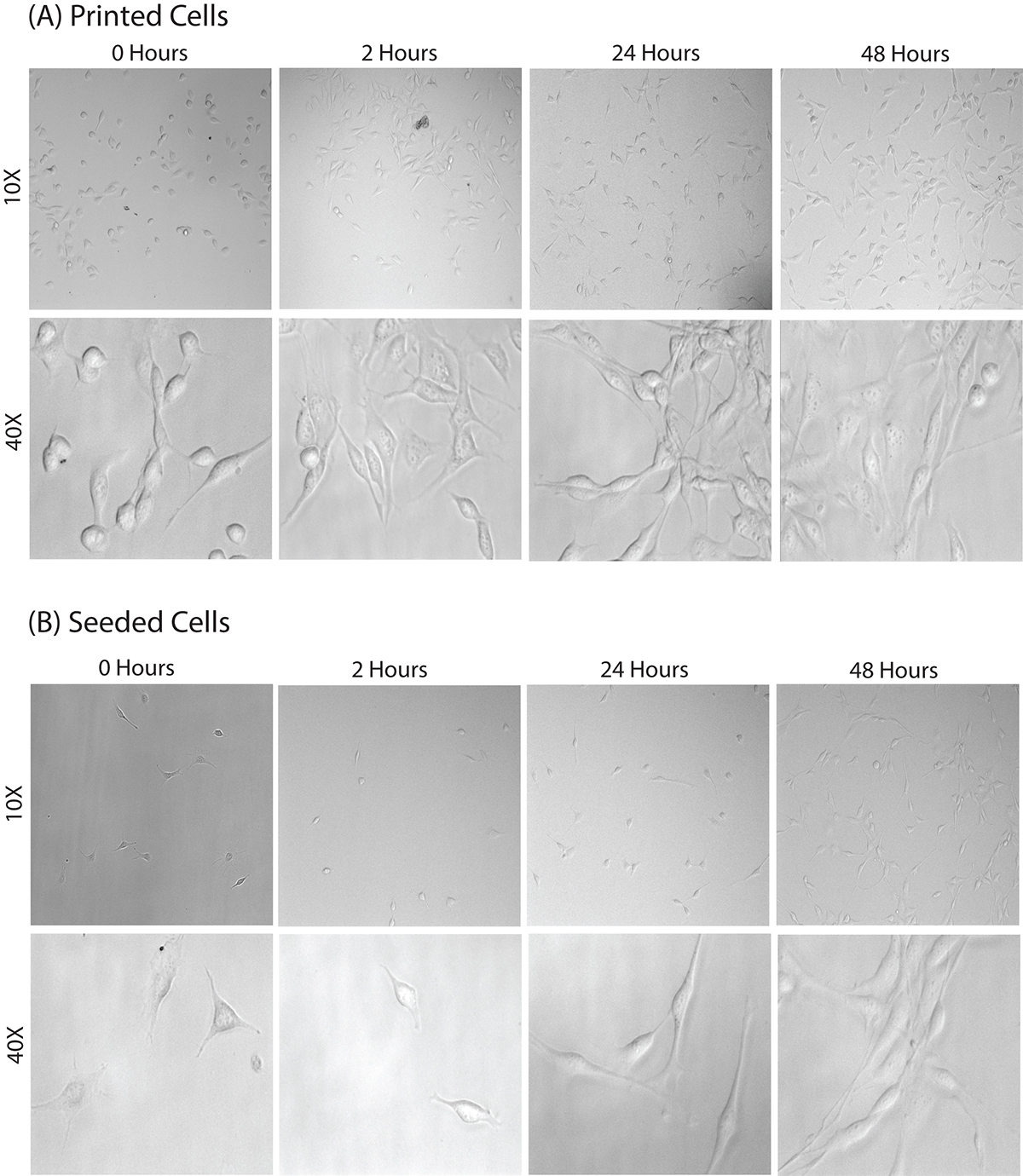
Figure 2. Images comparing morphology of seeded versus printed cells. In (A) 3T3 cells were microfluidically printed onto BSA and assessed at 0, 2, 24, and 48 hr. In (B) 3T3 cells were seeded instead of printed. The cell morphology is similar if not identical. Please click here to view a larger version of this figure.

Figure 3. Comparison of cell density and viability between cell printing and cell seeding at various time points including 0, 2, 24, and 48 hr. In (A) the cell density in cells per mm2 was assessed. In (B) the viability of the cells was determined. Please click here to view a larger version of this figure.
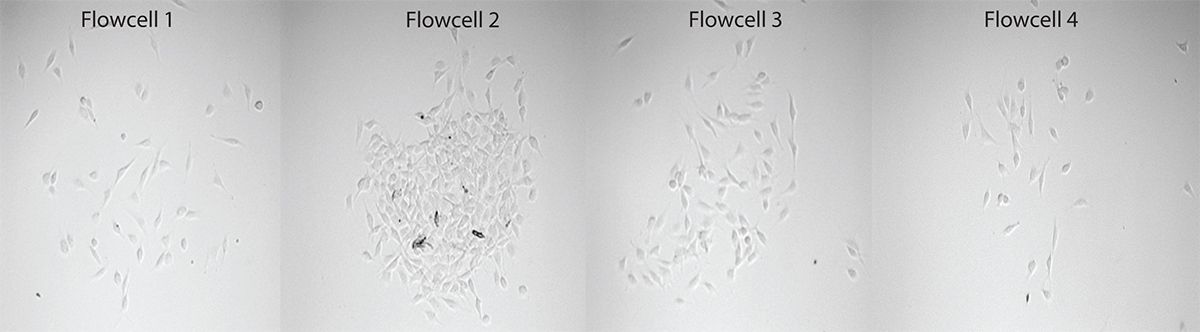
Figure 4. 10x magnification image of four flow cells of the cell microarray continuous flow microspotter in which NIH/3T3 cells were flowed through. Please click here to view a larger version of this figure.
Discussion
The 3D microfluidic printing technology described here is uniquely capable of microfluidically printing arrays of cells into a liquid filled well, i.e. a submerged surface. By printing onto submerged surfaces, cell microarrays can be produced that maintain the physiologically relevant cellular phenotype of cells as well as the ability to multiplex cells in the bottom of a single well Figure 4. The results of this study show that microfluidically printing cells results in cell attachment with comparable cell morphology and viability; however, the printed cells can be more densely attached in a defined location resulting in shorter overall study times. Shorter study times could result in significant cost savings for high throughput drug screening14.
Microfluidics has previously been used for cellular microarrays and high-throughput drug screening. Dozens of studies have detailed how flow based assays eliminate the static flow issues; unfortunately, these studies use 2D microfluidics (not to be confused with 2D and 3D tissue cultures), which limits density, multiplexing, and microscope integration for highly parallel analyses15–19. Digital microfluidics and droplet-based microfluidics have been proposed for these types of applications and can operate in highly-parallel, multiplexed configurations15,17,18, but they are limited to single or small cell groups and lose all 3-D, multi-cell type tissue structure during sample preparation. High throughput flow cytometry has been utilized for rapid cell screening; however, flow cytometry requires cells to remain in suspension, which is not ideal for screening adherent cell lines that, in vivo, are tethered to extracellular matrix20. Co-cultures also suffer from lack of true tissue representation19. In contrast, the 3D MFCA technology enables fully automated deposition of biomolecules and delivery of reagent to cell spots or intact tissue slices in a densely packed array. The proposed system would not only be able to generate arrays, but would do so in a “flowing” environment that allows for studies of shear stresses, delivery of different compounds, and growth conditions on these arrays and will lead to more predictive in vitro drug screening tools. A variety of applications can be envisioned allowing innovation to continue indefinitely. Such a tool enables novel cellular drug discovery and cytotoxicological assays in a multitude of critical research fields such as cancer, diabetes, inflammation, infections, and cardiovascular disease.
While the MFCA is the preferred method to deliver cells to a submerged surface, the challenge remains to attach the cell to the surface, especially for non-adherent cell lines. Future directions for this technology will focus on methods for cell attachment for the surface. Techniques for cell patterning on artificial surfaces currently exist and include: inkjet printing, microextrusion or filament plotting, DNA hybridization, and laser forward transfer21. Of these techniques, the attachment of cells using DNA hybridization technology possesses several unique and important features. First, the method of cell attachment is not dependent on receptors possessed by individual cells so both adherent and nonadherent cells are capable of being patterned using this method22. Next, the DNA cellular patterning method allows for the generation of complex cellular arrays comprising many cell types through the use of multiple surface-bound DNA sequences23. Future applications or directions will focus on using DNA hybridization technology for cell attachment19,24
Disclosures
The authors have nothing to disclose.
Acknowledgements
The authors would like to acknowledge Chris Morrow for technical assistance. Funding was provided by NIH SBIR (R43) grant 1R43GM101859-01 (MPI) GRANT10940803.
Materials
| Continuous Flow Microspotter | Wasatch Microfluidics | ||
| NIH/3T3 cells | ATCC | CRL-1658 | |
| Dubbleco's Modified Eagle Medium | Invitrogen | 11965-092 | base media for cells |
| HEPES buffer | Invitrogen | 15630-080 | cell media additive (control pH) |
| Sodium pyruvate | Invitrogen | 11360-070 | cell media additive |
| Penicillin-Streptomycin | Invitrogen | cell media additive | |
| Trypan blue | Invitrogen | 15250-061 | stain cell sfor counting |
| Haemocytometer | Fisher | 267110 | cell chamber to count cells |
| Nikon Eclipse TS100 | Nikon | Used to check on cells | |
| Nikon Eclipse TE2000-U | Nikon | Used for collecting images | |
| Phosphate Buffered Saline (with calcium and magnesium) | Invitrogen | 14040-133 | rinsing cells before passaging and before staining with PI |
| TrypLE Express | Invitrogen | A12177-01 | used to remove cells from surface |
References
- Fernandes, T. G., Diogo, M. M., Clark, D. S., Dordick, J. S., Cabral, J. High-throughput cellular microarray platforms: applications in drug discovery, toxicology and stem cell research. Trends in Biotechnology. 27, 342-349 (2009).
- Michelini, E., Cevenini, L., Mezzanotte, L., Coppa, A., Roda, A. Cell-based assays: fuelling drug discovery. Anal Bioanal Chem. 398, 227-238 (2010).
- Gao, D., et al. Recent developments in microfluidic devices for in vitro cell culture for cell-biology research. TrAC Trends in Analytical Chemistry. 35, 150-164 (2012).
- Geysen, H. M., Schoenen, F., Wagner, D., Wagner, R. Combinatorial compound libraries for drug discovery: an ongoing challenge. Nature Reviews Drug Discovery. 2, 222-230 (2003).
- Xu, Y., Shi, Y., Ding, S. A chemical approach to stem-cell biology and regenerative medicine. Nature. 453, 338-344 (2008).
- Khademhosseini, A., Langer, R., Borenstein, J., Vacanti, J. P. Microscale technologies for tissue engineering and biology. Proceedings of the National Academy of Sciences of the United States of America. 103, 2480-2487 (2006).
- Bhadriraju, K., Chen, C. S. Engineering cellular microenvironments to improve cell-based drug testing. Drug Discovery Today. 7, 612-620 (2002).
- Castel, D., Pitaval, A., Debily, M. -. A., Gidrol, X. Cell microarrays in drug discovery. Drug Discov. Today. 11, 616-622 (2006).
- Tsui, J. H., Lee, W., Pun, S. H., Kim, J., Kim, D. -. H. Microfluidics-assisted in vitro drug screening and carrier production. Advanced Drug Delivery Reviews. , (2013).
- Gottwald, E., et al. A chip-based platform for the in vitro generation of tissues in three-dimensional organization. Lab on a Chip. 7, 777-785 (2007).
- Liu, R., Lee, A. P. . Integrated Biochips for DNA Analysis. , (2008).
- Natarajan, S., et al. Continuous-flow microfluidic printing of proteins for array-based applications including surface plasmon resonance imaging. Analytical Biochemistry. 373, 141-146 (2008).
- Natarajan, S., Hatch, A., Myszka, D. G., Gale, B. K. Optimal Conditions for Protein Array Deposition Using Continuous Flow. Anal. Chem. 80, 8561-8567 (2008).
- Keighley, W. The need for high throughput kinetics early in the drug discovery process. , (2011).
- Xu, F., et al. A three-dimensional in vitro ovarian cancer coculture model using a high-throughput cell patterning platform. Biotechnol J. 6, 204-212 (2011).
- Tuckwell, D. S., Weston, S. A., Humphries, M. J. Integrins: a review of their structure and mechanisms of ligand binding. Symposia of the Society for Experimental Biology. 47, 107 (1993).
- Roberts, C., et al. Using mixed self-assembled monolayers presenting RGD and (EG) 3OH groups to characterize long-term attachment of bovine capillary endothelial cells to surfaces. Journal of the American Chemical Society. 120, 6548-6555 (1998).
- Chandra, R. A., Douglas, E. S., Mathies, R. A., Bertozzi, C. R., Francis, M. B. Programmable cell adhesion encoded by DNA hybridization. Angewandte Chemie. 118, 910-915 (2006).
- Hsiao, S. C., et al. Direct cell surface modification with DNA for the capture of primary cells and the investigation of myotube formation on defined patterns. Langmuir. 25, 6985-6991 (2009).
- Black, C. B., Duensing, T. D., Trinkle, L. S., Dunlay, R. T. Cell-Based Screening Using High-Throughput Flow Cytometry. ASSAY and Drug Development Technologies. 9, 13-20 (2011).
- Derby, B. Bioprinting: inkjet printing proteins and hybrid cell-containing materials and structures. J. Mater. Chem. 18, 5717-5721 (2008).
- Onoe, H., et al. Cellular Microfabrication: Observing Intercellular Interactions Using Lithographically-Defined DNA Capture Sequences. Langmuir. 28, 8120-8126 (2012).
- Hsiao, S. C., et al. DNA-Coated AFM Cantilevers for the Investigation of Cell Adhesion and the Patterning of Live Cells. Angewandte Chemie International Edition. 47, 8473-8477 (2008).

