Linear Amplification Mediated PCR – Localization of Genetic Elements and Characterization of Unknown Flanking DNA
Summary
Linear-amplification mediated (LAM)-PCR is a method developed to identify the exact positions of integrating viral vectors in the genome. The technique has evolved to be the superior method to study clonal dynamics in gene therapy patients, biosafety of novel vector technologies, T-cell diversity, cancer stem cell models, etc.
Abstract
Linear-amplification mediated PCR (LAM-PCR) has been developed to study hematopoiesis in gene corrected cells of patients treated by gene therapy with integrating vector systems. Due to the stable integration of retroviral vectors, integration sites can be used to study the clonal fate of individual cells and their progeny. LAM- PCR for the first time provided evidence that leukemia in gene therapy treated patients originated from provirus induced overexpression of a neighboring proto-oncogene. The high sensitivity and specificity of LAM-PCR compared to existing methods like inverse PCR and ligation mediated (LM)-PCR is achieved by an initial preamplification step (linear PCR of 100 cycles) using biotinylated vector specific primers which allow subsequent reaction steps to be carried out on solid phase (magnetic beads). LAM-PCR is currently the most sensitive method available to identify unknown DNA which is located in the proximity of known DNA. Recently, a variant of LAM-PCR has been developed that circumvents restriction digest thus abrogating retrieval bias of integration sites and enables a comprehensive analysis of provirus locations in host genomes. The following protocol explains step-by-step the amplification of both 3’- and 5’- sequences adjacent to the integrated lentiviral vector.
Introduction
Linear-amplification mediated PCR (LAM-PCR) allows identifying and characterizing unknown flanking DNA adjacent to known DNA of any origin. More specifically, LAM-PCR has been developed to localize viral vector integration sites (IS) within the host genome1,2. Genetic elements like retroviruses or transposons integrate their genome into the host genome in a (semi-) random manner3-6. In many cases it is decisive to know exactly the position where these vectors integrated. LAM-PCR has been proven to be superior to alternative techniques like ligation-mediated PCR7 and its variants or inverse PCR8. The sensitivity and robustness of this method arises from the initial preamplification of the vector-genome junctions and magnetic selection of amplified PCR products. Like the alternative methods mentioned, LAM-PCR relies on the use of restriction enzymes, introducing a bias into retrieval capacity of IS9-11. Thus, only a subset of the IS repertoire (the integrome) can be detected in one reaction. This bias is minimized by the parallel analysis of a given sample using optimal combinations of restriction enzymes9. Recently, a variant of the technology termed non-restrictive LAM-PCR (nrLAM-PCR) has been developed that circumvents the use of restriction enzymes and allows unbiased genome-wide analysis of a sample in a single reaction9,12.
In the past, LAM-PCR has been used to identify the causative retroviral IS giving rise to leukemia in a few patients in clinical gene therapy trials13-15. Since then, LAM-PCR has been adapted to identify IS from other integrating vectors (lentiviral vectors, transposons) and also to identify integration patterns of passively integrating vectors like adeno-associated vectors (AAV) or integrase-defective lentiviral vectors (IDLV)16-21. Applications of LAM-PCR are wide spread: traditionally, the technique is widely used to study the clonal composition of gene modified cells in patients that have undergone gene therapy or to assess the biosafety of novel vector systems by unraveling their integration behavior15,16,22-24. Recently, LAM-PCR enabled determining specificity and off-target activity of designer nucleases by an IDLV trapping assay25.
Moreover, LAM-PCR allows to easily follow the fate of a transduced cell over time in an organism. This allows to identify proto-oncogenes as well as tumor suppressor genes and also to study hematopoiesis or cancer stem cell biology26-28. Last but not least, LAM-PCR was adapted to study T-cell receptor diversity in humans29 (and unpublished data).
The intrinsic power of the technology is reinforced by linking the method to deep sequencing technologies that allow characterizing millions of unknown flanking DNA with single nucleotide resolution in whole genomes. In the following protocol, we describe step-by step the amplification and identification of flanking unknown DNA exemplarily to identify lentiviral vector IS. Oligonucleotides used in the protocol are listed in Table 1. Extracted DNA or cDNA of any source can be used as DNA template for LAM-PCR and nrLAM-PCR.
Protocol
1. Preparation of Linker Cassettes (LC)
- Mix 40 µl of LC1 oligonucleotide (Table 1), 40 µl of LC2 oligonucleotide (Table 1, with proper restriction enzyme overhang), 110 µl Tris-HCl (100 mM, pH 7.5), and 10 µl 250 mM MgCl2.
- Incubate at 95 °C for 5min and let the reaction cool down slowly to room temperature. Add 300 µl H2O and concentrate dsLinker-DNA on a centrifugation filter. Add 80 µl H2O to the eluate and aliquot 10 µl of prepared linker cassette in 0.2 PCR tubes.
2. Preamplification of Vector Genome Junctions
- For every sample to be analyzed prepare a 50 µl PCR reaction.
- Determine concentration of the DNA samples. Pipette x µl (1-1,000 ng for LAM/100-1,000 ng for nrLAM) of DNA into a 0.2 ml PCR tube. Volume of DNA should be equal in each sample and in the range of 0.5 to 25 µl.
- Prepare PCR master mix as described in Table 2. Mix (50 – x) µl of the master mix with each DNA sample in 0.2 ml PCR tubes.
- Preamplify vector genome junctions using PCR conditions exemplified in Table 2. After completion of PCR add 0.5 µl Taq Polymerase to each PCR tube and rerun PCR program. PCR products can be stored at 4 °C for up to 4 days or long term at -20 °C.
3. Magnetic Separation of PCR Product
- Preparation of Magnetic Beads
- Pipette 20 µl (200 µg) of streptavidin coated magnetic beads into a 1.5 ml tube and expose for 1 min on the magnetic particle separator (MPS) at room temperature. Discard supernatant.
- Remove tube from MPS and resuspend magnetic beads in 40 µl PSB/ 0.1% BSA (pH 7.5). Expose to MPS for 1 min and discard supernatant. Repeat this step once.
- Wash beads with 20 µl of 3 M LiCl solution (3 M LiCl, 10 mM Tris-HCl, 1 mM EDTA) expose to MPS for 1 min and discard supernatant. Resuspend beads in 50 µl of 6 M LiCl solution (6 M LiCl, 10 mM Tris-HCl, 1 mM EDTA).
- Mix entire PCR reaction from step 2.2 with 50 µl of prepared magnetic beads. Incubate on a horizontal shaker (300 rpm) at least for 2 hr at room temperature. This step allows binding of biotinylated PCR product to streptavidin coated beads (DNA-Bead complex). DNA bead complex can be incubated on the shaker overnight or stored at 4 °C for up to 4 days.
- Expose DNA-Bead complex to MPS for 1 min at room temperature, remove supernatant, and resuspend DNA-Bead complex in 100 µl H2O. Immediately proceed with step 4 for LAM or step 5 for nrLAM.
4. LAM-Procedure
- Double Strand DNA (dsDNA) Synthesis (LAM only)
- Expose DNA-Bead complex from step 3.3 to MPS for 1 min and discard supernatant. Add 8.25 µl of H2O, 1 µl 10x hexanucleotide buffer, 0.25 µl dNTPs (10 mM) and 0.5 µl (2 U) Klenow polymerase. Incubate at 37 °C for 1 hr.
- Add 90 µl of H2O and expose to MPS for 1 min. Discard supernatant and resuspend DNA-Bead complex in 100 µl H2O.
- Restriction Digest
- Expose DNA-Bead complex to MPS for 1 min and discard supernatant. Add 8.5 µl H2O, 1 µl 10x restriction enzyme buffer and 0.5 µl of restriction enzyme and incubate reaction for 1 hr. Repeat step 4.1.2.
NOTE: Incubate reaction at the temperature recommended by the manufacturer of the restriction enzyme. Make sure that there is no restriction site present within or downstream of the primer binding site used for preamplification in the DNA of interest. For choice of suitable restriction enzymes/restriction enzyme combinations refer to 9.
- Expose DNA-Bead complex to MPS for 1 min and discard supernatant. Add 8.5 µl H2O, 1 µl 10x restriction enzyme buffer and 0.5 µl of restriction enzyme and incubate reaction for 1 hr. Repeat step 4.1.2.
- Ligation of ds Linker (LK)
- Expose DNA-Bead complex to MPS for 1 min and discard supernatant. Add 5 µl H2O, 1 µl 10x FastLink buffer, 1 µl ATP (10 mM), 2 µl Linker cassette from step 1.2, and 1 µl Fast-Link DNA Ligase (2 U/µl). Incubate at room temperature for 5 min. Repeat step 4.1.2.
- Denaturation of Synthesized dsDNA
- Expose DNA-Bead complex to MPS for 1 min and discard supernatant. Resuspend DNA-Bead complex in 5 µl 0.1 N NaOH. Incubate 5 min at room temperature on a horizontal shaker.
- Expose DNA-Bead complex to MPS for 1 min and collect preamplified vector-genome junction containing supernatant in new 1.5 ml tube. Immediately proceed with step 6 or store supernatant at -20 °C.
5. nrLAM-Procedure
- Ligation of single stranded linker (ssLC) (nrLAM only)
- Expose DNA-Bead complex from step 3.3 to MPS for 1 min and discard supernatant. Add 6.5 µl H2O, 1 µl CircLigase 10x Reaction Buffer, 0.5 µl MnCl2 (50 mM), 0.5 µl ATP (1 mM), 1 µl ssLinker oligonucleotide and 0.5 µl CircLigase (100 U/µl). Incubate at 60 °C for 1 hr.
- Add 90 µl of H2O and expose to MPS for 1 min. Discard supernatant and wash DNA-Bead complex in 100 µl H2O. Again expose to MPS for 1 min, discard supernatant and resuspend DNA-Bead complex in 10 µl H2O.
6. Exponential Amplification I
- For every sample to be analyzed prepare a 50 µl PCR reaction.
- Pipette 2 µl template DNA (from step 4.4.2 (LAM) or 5.1.2 (nrLAM)) into a 0.2 ml PCR tube.
- Prepare PCR master mix as described in Table 3. Add 48 µl of master mix to each sample from step 6.1.1 and amplify vector genome junctions by PCR conditions exemplified in Table 3. PCR products can be stored at 4 °C for up to 4 days or long term at -20 °C.
7. Magnetic Separation of PCR Product
- Prepare magnetic beads as exemplified in steps 3.1.1 – 3.1.3. Resuspend beads in 20 µl (for LAM) or 50 µl (for nrLAM) of 6 M LiCl solution (6 M LiCl, 10 mM Tris-HCl, 1 mM EDTA). Mix 20 µl (LAM) or 50 µl (nrLAM) of PCR reaction from step 6.1.2 with prepared magnetic beads (step 7.1) and incubate on a horizontal shaker (300 rpm) for 2 hr at room temperature. DNA bead complex can be incubated on the shaker overnight or stored at 4 °C for up to 4 days.
- Expose DNA-Bead complex to MPS for 1 min, remove supernatant and resuspend DNA-Bead complex in 100 µl H2O. Expose DNA-Bead complex to MPS for 1 min and discard supernatant.
- Resuspend DNA-Bead complex in 20 µl (LAM) or 5 µl (nrLAM) 0.1 N NaOH. Incubate for 10 min at room temperature on a horizontal shaker, expose to MPS for 1 min and collect amplified DNA containing supernatant in new 1.5 ml tube. Immediately proceed with step 8.1 or store supernatant at -20 °C.
8. Exponential Amplification II
- For every sample to be analyzed prepare a 50 µl PCR reaction.
- Pipette 2 µl template DNA (from step 7.3) into a 0.2 ml PCR tube.
- Prepare PCR master mix as described in Table 4. Add 48 µl of master mix to each sample from step 8.1.1 and amplify vector genome junctions by PCR conditions exemplified in Table 4. PCR products can be stored at 4 °C for up to 4 days or long term at -20 °C.
- To visualize (nr)LAM-PCR products, load 10 µl of the PCR product from step 8.1.2 on a 2% agarose gel. If bands are visible, analyze 10 µl of the LAM-PCR product on high-resolution gel. CAUTION: Ethidium bromide is mutagenic. Work very carefully and always wear appropriate gloves.
9. Preparation for High-throughput Sequencing
- Purification of (nr)LAM-PCR products
- Mix 40 µl PCR-Product from step 8.1.2 with 44 µl of room temperature AMPure XP Magnetic Beads. Incubate 5 min at room temperature and expose to MPS for additional 2 min.
- Discard supernatant and wash twice with 200 µl 70% EtOH on the MPS.
- Discard supernatant and resuspend DNA-Bead complex in 30 µl H2O. Incubate 1 min and transfer supernatant to fresh 0.2 ml tube. Determine concentration of purified DNA.
- Fusionprimer PCR to add sequencing specific adaptors.
- For every sample to be analyzed prepare a 50 µl PCR reaction.
- Pipette x µl (40 ng) of DNA into a 0.2 ml PCR tube. Volume of DNA should be equal in each sample and in the range of 0.5 to 25 µl.
- Prepare PCR master mix as described in table 5. Add (50 – x) µl of master mix to each sample from step 9.2.2 and introduce Sequencing adaptors to (nr)LAM-PCR products by PCR conditions exemplified in Table 5. PCR products can be stored at 4 °C for up to 4 days or long term at -20 °C.
- To visualize Fusionprimer-PCR products load 10 µl of the PCR product from step 9.2.3 on a 2% agarose gel. Purify remaining PCR product as described in steps 9.1.1 – 9.1.3. Analyze 1 µl purified PCR product from step 9.4 on an automated high-resolution electrophoresis device to accurately quantify concentration and fragment size of Fusionprimer-PCR products.
Representative Results
LAM-PCR results in amplification of vector genome junctions with a defined fragment size for each junction. The size of individual PCR fragments depends on the distance between the location of the known DNA in the genome and the closest restriction enzyme recognition site. This allows visualizing the diversity of amplified junctions in analyzed samples by gel electrophoresis, e.g., if only single (monoclonal), several (oligoclonal), or multiple (polyclonal) bands are present on the gel. The results of LAM-PCR are best viewed by high resolution electrophoresis gels (Figure 2A) but can also be visualized on 2% agarose gels (Figure 2B). nrLAM-PCR results in PCR fragments of various length for each individual junction. Thus monoclonal, oligoclonal or polyclonal samples appear as a smear by electrophoresis and cannot be distinguished visually. Visualizing the nrLAM-PCR product on 2% agarose gel is sufficient to determine success of the protocol (Figure 2C). After sequencing the recovered genomic DNA can be aligned to the respective host genome to identify exact positions of the location of the vector (Figure 3A). Annotation of the genome allows to analyze the IS repertoire for different vector specific features like preference for integration into gene coding regions (Figure 3B) or close to transcription start sites (Figure 3C).
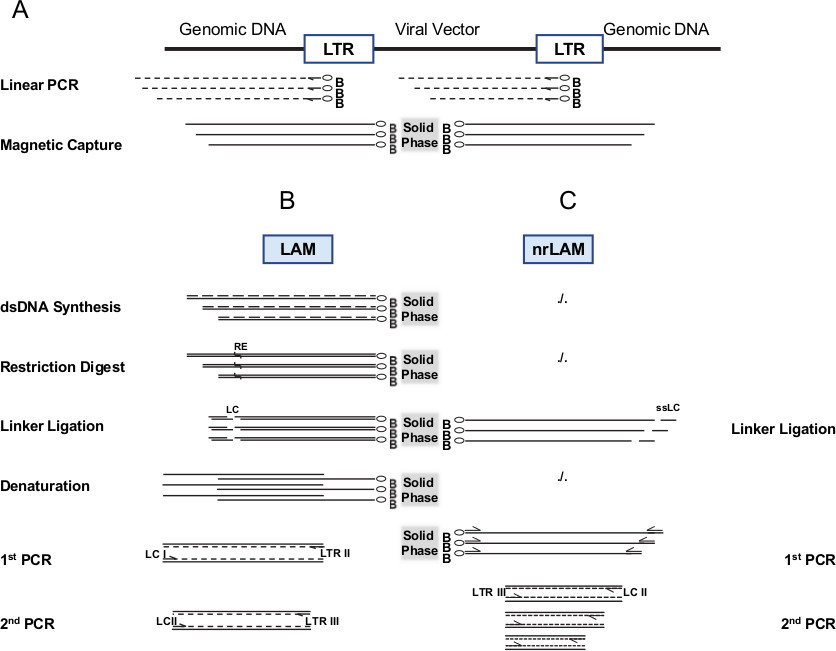
Figure 1. Schematic outline of LAM-PCR and nrLAM-PCR. A) Both methods start with an initial preamplification of vector genome junctions using biotinylated primers hybridizing close to the end of the known DNA sequence (here long terminal repeat (LTR) of a retroviral vector). Preamplification results in biotinylated ssDNA of different size for identical or different vector genome junctions. Biotinylated ssDNA is captured on magnetic particles. B) For LAM PCR, enzymatic reaction steps composed of dsDNA synthesis, restriction digest and ligation of a known linker DNA generate products of different sizes with known sequences on both ends of the product. Due to restriction length polymorphism each amplified junction has a characteristic length. After denaturation the LAM-PCR product is amplified by nested PCR with linker and vector specific primers. C) For nrLAM a ssDNA linker sequence is directly ligated to the unknown end of the preamplified ssDNA from A) allowing exponential amplification by nested PCR with linker and vector specific primers. This figure has been modified from 2,12. Please click here to view a larger version of this figure.

Figure 2. Representative results of LAM- PCR and nrLAM-PCR. A, B) LAM-PCR analysis of isolated DNA from peripheral blood of gene therapy treated patients. The number of bands on the gel corresponds to the number of IS present in the sample. High-resolution gels (B) are better suited to visualize clonality of analyzed samples than 2% agarose gels (A). C) nrLAM-PCR analysis of lentiviral vector transduced single cell clones or bulk cells. Independent of the number of amplified insertion sites a smear is seen on the gel after electrophoresis. M, 100 bp ladder; MC, monoclonal; OC, oligoclonal; PC, polyclonal. This figure has been modified from 2,9.

Figure 3. Representative examples for IS analysis by LAM-PCR and subsequent high-throughput sequencing. IS distribution in two patients from gammaretroviral (blue) or lentiviral (green) clinical gene therapy trials. After sequencing and mapping of LAM-PCR products to the respective genome IS can be evaluated e.g.: A) Genome-wide distribution of IS. B) Difference according to the preference for insertion into gene coding regions between gammaretroviral and lentiviral vectors and C) preference for insertion close to transcription start sites. Please click here to view a larger version of this figure.
| Purpose | Name | Sequence (5'-3') |
| LK-universal | LC1 | GACCCGGGAGATCTGAATTCAGTGGCACAG CAGTTAGG |
| LK-AATT | LC2 (AATT) | AATTCCTAACTGCTGTGCCACTGAATTCA GATC |
| LK-CG | LC2 (CG) | CGCCTAACTGCTGTGCCACTGAATTCAGATC |
| LK-TA | LC2 (TA) | TACCTAACTGCTGTGCCACTGAAATCAGATC |
| LK-nrLAM-PCR | ssLC | (P)CCTAACTGCTGTGCCACTGAATTCAGATC TCCCGGGTddC |
| Preamplification | LTR-I (3'-direction) | (B)AGTAGTGTGTGCCCGTCTGT |
| LTR-I (5'-direction) | (B)TTAGCCAGAGAGCTCCCAGG | |
| Exponential amplification I | LTR-II (3'-direction) | (B)GTGTGACTCTGGTAACTAGAG |
| LTR-II (5'-direction) | (B)GATCTGGTCTAACCAGAGAG | |
| LC-I | GACCCGGGAGATCTGAATTC | |
| Exponential amplification II | LTR-III (3'-direction) | GATCCCTCAGACCCTTTTAGTC |
| LTR-III (5'-direction) | CCCAGTACAAGCAAAAAGCAG | |
| LC-II | GATCTGAATTCAGTGGCACAG |
Table 1. Oligonucleotides for LAM- and nrLAM-PCR to amplify lentiviral IS. ssLC is phosphorylated at the 5’-end (P) and has at 3’ didesoxycytidin (ddC) to avoid multimerization of the ssLC during ligation. In general, (nr)LAM-PCR primers should consist of 18-25 nucleotides and should not align to the host genome. Primers for preamplification should be placed as close as possible (≤120 bp) to the 5′ or 3′ end of the vector. Two additional primers for Exponential PCR I and II need to be placed between the primer used for preamplification and the vector end. Primers for preamplification and Exponential PCR I need to be 5’-phosphorylated (P).
| Reagent | Volume (µl) | Concentration | PCR Parameters | Temperature | Time | |
| H2O | 43 – x | Initial denaturation | 95 °C | 5 min | ||
| Buffer | 5 | 10 x | Denaturation | 95 °C | 45 sec | |
| dNTP | 1 | 10 mM (LAM); 0.5 µM (nrLAM) | Annealing | 60 °C | 45 sec | 2 x 50 Cycles |
| LTR-I | 0.5 | 0.17 µM | Elongation | 72 °C | 60 sec (LAM); 10 sec (nrLAM) | |
| Taq Polymerase | 0.5 | 2.5 U/µl | Final Elongation | 72 °C | 5 min (only LAM) |
Table 2. PCR-Conditions for preamplification of vector genome junctions (step 2). Columns 1-3 show the PCR reagents used for amplification of a single DNA sample. Columns 4-6 exemplify the PCR program to preamplify vector genome junctions.
| Reagent | Volume (µl) | Concentration | PCR Parameters | Temperature | Time | |
| H2O | 40.5 | Initial denaturation | 95 °C | 5 min | ||
| Buffer | 5 | 10 x | Denaturation | 95 °C | 45 sec | |
| dNTP | 1 | 10 mM | Annealing | 60 °C | 45 sec | 35 Cycles |
| LTR-II | 0.5 | 16.7 µM | Elongation | 72 °C | 60 sec (LAM); 5 sec (nrLAM) | |
| LC-I | 0.5 | 16.7 µM | Final Elongation | 72 °C | 5 min (only LAM) | |
| Taq Polymerase | 0.5 | 2.5 U/µl |
Table 3. PCR-Conditions for exponential Amplification I (step 6). Columns 1-3 show the PCR reagents used for exponential amplification of a single DNA sample. Columns 4-6 exemplify the PCR program used to exponentially amplify one sample after Ligation of linker sequence.
| Reagent | Volume (µl) | Concentration | PCR Parameters | Temperature | Time | |
| H2O | 40.5 | Initial denaturation | 95 °C | 5 min | ||
| Buffer | 5 | 10 x | Denaturation | 95 °C | 45 sec | |
| dNTP | 1 | 10 mM | Annealing | 60 °C | 45 sec | 35 Cycles |
| LTR-III | 0.5 | 16.7 µM | Elongation | 72 °C | 60 sec (LAM); 5 sec (nrLAM) | |
| LC-II | 0.5 | 16.7 µM | Final Elongation | 72 °C | 5 min | |
| Taq Polymerase | 0.5 | 2.5 U/µl |
Table 4. PCR-Conditions for exponential Amplification I (step 8). Columns 1-3 show the PCR reagents used for nested exponential amplification of a single sample. Columns 4-6 exemplify the PCR program used for nested exponential amplification of vector genome junctions from one sample.
| Reagent | Volume (µl) | Concentration | PCR Parameters | Temperature | Time | |
| H2O | 42.5 – x | Initial denaturation | 95 °C | 2 min | ||
| Buffer | 5 | 10 x | Denaturation | 95 °C | 45 sec | |
| dNTP | 1 | 10 mM | Annealing | 58 °C | 45 sec | 12 Cycles |
| Fusionprimer A | 0.5 | 10 µM | Elongation | 72 °C | 60 sec | |
| Fusionprimer B | 0.5 | 10 µM | Final Elongation | 72 °C | 5 min | |
| Taq Polymerase | 0.5 | 2.5 U/µl |
Table 5. PCR-Conditions for Fusionprimer-PCR (step 9.2). Columns 1-3 show the PCR reagents used for introduction of sequencing adaptors to (nr)LAM-PCR products. Columns 4-6 exemplify the PCR program used for Fusionprimer-PCR.
Discussion
The LAM-PCR technique allows identifying unknown DNA sequences that flank a known DNA region. Because of the high sensitivity resulting from preamplification of the junctions with specific primers hybridizing in the known DNA sequence, it is possible to amplify and detect even rare junctions down to the single cell level. Contrary, in a polyclonal situation LAM-PCR is able to amplify thousands of different junctions in one single reaction.
However, due to the use of restriction enzymes only a subfraction of the integrome can be analyzed by LAM-PCR for the presence of junctions with every particular restriction enzyme. Thus, repeated analysis of the same sample with different enzymes is recommended9. If no LAM-PCR amplicons are present on the gel, most likely the distance between the location of the known DNA fragment and the closest recognition site of the chosen restriction enzyme is too large to result in LAM-PCR products9. In this case other enzymes should be used to amplify the junction.
nrLAM is independent of the use of restriction enzymes and therefore represents a highly valuable method to comprehensively characterize sequences flanking a known DNA sequence. Omitting restriction digest from the protocol results in the loss of specific restriction fragment length polymorphism characterizing each amplified junction. Instead every amplified junction is represented by PCR products of various sizes resulting in a smear on the gel after electrophoresis, independent of the diversity of amplified junctions.
Both LAM- and nrLAM-PCR products are perfectly suited for downstream high-throughput sequencing. High-throughput sequencing of (nr)LAM-PCR products and mapping of retrieved raw sequences to the corresponding genome allows characterizing unknown flanking DNA or identifying the exact localization of vector-genome junctions30. By introducing barcode sequences into the fusionprimers several hundreds of LAM-and nrLAM-PCR products can be sequenced in one sequencing run30.
Due to high sensitivity, LAM-PCR is prone to contamination if executed inattentively. Thus, a PCR-grade environment and special attention to clean handling of the protocol is of utmost importance to successfully amplify the unknown flanking DNA without contaminating samples. Therefore, including untransduced genomic DNA and a water control for every PCR reaction as negative controls into the LAM-PCR protocol is strongly recommended. If control samples indicate that cross-contamination occurred during the protocol, the products from every pause point can be used to repeat parts of the protocol. When bands are still present, it is recommended to discard all reagents (e.g., primers, dNTPs, polymerases, etc.) and repeating the (nr)LAM-PCR protocol with new aliquots.
Disclosures
The authors have nothing to disclose.
Acknowledgements
Funding was provided by the Deutsche Forschungsgemeinschaft (SPP1230, grant of the Tumor Center Heidelberg/Mannheim), by the Bundesministerium für Bildung und Forschung (iGene), by the VIth + VIIth Framework Programs of the European Commission (CONSERT, CLINIGENE and PERSIST). We thank Ina Kutschera for demonstrating the protocol technique in the video.
Materials
| Name of Material/ Equipment | Company | Catalog Number | Comments/Description |
| Taq DNA Polymerase | Genaxxon Bioscience GmbH | M3001.5000 | Alternative Taq Polymerases may be used |
| PCR Buffer | Qiagen | 201203 | Use of this buffer is recommended |
| dNTP-Mixture | Genaxxon Bioscience GmbH | M3015.4020 | or any other dNTPs |
| Oligonucleotides (Primers) | MWG Biotech | HPLC purified | |
| Dynabeads M-280 Streptavidin | Invitrogen | 11206D | |
| PBS | Gibco | 14190-086 | 0.1 % wt/vol BSA |
| 6M LiCl | Roth | 3739.1 | 10 mM Tris-HCl (pH 7.5)/1 mM EDTA |
| Tris-HCl, pH 7.5 | USB Corporation | 22637 | or any other supplier |
| EDTA | Applichem | A1103,0250 | or any other supplier |
| Klenow Polymerase | Roche Diagnostics | 10104523001 | |
| Hexanucleotide mixture | Roche Diagnostics | 11277081001 | |
| Restriction endonuclease | NEB | or any other supplier | |
| Fast-Link DNA ligation kit | Epicentre Biotechnologies | LK11025 | |
| CircLigase ssDNA Ligase Kit | Epicentre Biotechnologies | CL4111K | |
| NaOH | Sigma-Aldrich | 72068 | or any other supplier |
| Agarose LE | Roche Diagnostics | 11685660001 | or any other supplier |
| TBE buffer | Amresco | 0658 | or any other supplier |
| Ethidium bromide | Applichem | A2273,0005 | Ethidium bromide is mutagenic |
| 100 bp DNA Ladder | Invitrogen | 15628-050 | or any other DNA ladder |
| 20 mM NaCl | Sigma-Aldrich | 71393-1L | or any other supplier |
| Magna-Sep Magnetic Particle Separator | Life Technologies | K158501 | for use with 1.5 ml Tubes |
| Magna-Sep Magnetic Particle Separator | Life Technologies | K158696 | for use with 96 well plates |
| Amicon Ultra-0.5, Ultracel-30 membrane | Millipore | UFC503096 | |
| PerfectBlue Gelsystem Midi S | PeqLab | 40-1515 | or other electrophoresis system |
| TProfessional 96 | Biometra | 050-551 | or other Thermocycler for 96-well plates |
| Orbital shaker KS 260 basic | IKA | 2980200 | or other horizontal shaker |
| PCR softtubes 0.2 ml | Biozym Scientific GmbH | 711082 | or other 0.2 ml PCR tubes |
| 1.5 ml tubes | Eppendorf | 12682 | or other 1.5 ml tubes |
| Gel documentation system | PeqLab | or any other gel documentation system | |
| Nanodrop ND-1000 spectrophotometer | Thermo Scientific | ND-1000 | |
| Spreadex EL1200 precast gel | Elchrom Scientific | 3497 | |
| Submerged gel electrophoresis apparatus SEA 2000 | Elchrom Scientific | 2001E | |
| 2100 Electrophoresis Bioanalyzer | Agilent Technologies | G2939AA |
References
- Schmidt, M., et al. Detection and direct genomic sequencing of multiple rare unknown flanking DNA in highly complex samples. Hum Gene Ther. 12, 743-749 (2001).
- Schmidt, M., et al. High-resolution insertion-site analysis by linear amplification-mediated PCR (LAM-PCR). Nat Methods. 4, 1051-1057 (2007).
- Schroeder, A. R., et al. HIV-1 integration in the human genome favors active genes and local hotspots. Cell. 110, 521-529 (2002).
- Wu, X., Li, Y., Crise, B., Burgess, S. M. Transcription start regions in the human genome are favored targets for MLV integration. Science. 300, 1749-1751 (2003).
- Vigdal, T. J., Kaufman, C. D., Izsvak, Z., Voytas, D. F., Ivics, Z. Common physical properties of DNA affecting target site selection of sleeping beauty and other Tc1/mariner transposable elements. J Mol Biol. 323, 441-452 (2002).
- Lewinski, M. K., et al. Retroviral DNA integration: viral and cellular determinants of target-site selection. PLoS Pathog. 2, (2006).
- Mueller, P. R., Wold, B. In vivo footprinting of a muscle specific enhancer by ligation mediated PCR. Science. 246, 780-786 (1989).
- Silver, J., Keerikatte, V. Novel use of polymerase chain reaction to amplify cellular DNA adjacent to an integrated provirus. Journal of virology. 63, 1924-1928 (1989).
- Gabriel, R., et al. Comprehensive genomic access to vector integration in clinical gene therapy. Nature medicine. 15, 1431-1436 (2009).
- Harkey, M. A., et al. Multiarm high-throughput integration site detection: limitations of LAM-PCR technology and optimization for clonal analysis. Stem Cells Dev. 16, 381-392 (2007).
- Wang, G. P., et al. DNA bar coding and pyrosequencing to analyze adverse events in therapeutic gene transfer. Nucleic acids research. 36, (2008).
- Paruzynski, A., et al. Genome-wide high-throughput integrome analyses by nrLAM-PCR and next-generation sequencing. Nat. Protocols. 5, 1379-1395 (2010).
- Hacein-Bey-Abina, S., et al. Insertional oncogenesis in 4 patients after retrovirus-mediated gene therapy of SCID-X1. J Clin Invest. , 3132-3142 (2008).
- Hacein-Bey-Abina, S., et al. LMO2-associated clonal T cell proliferation in two patients after gene therapy for SCID-X1. Science. 302, 415-419 (2003).
- Howe, S. J., et al. Insertional mutagenesis combined with acquired somatic mutations causes leukemogenesis following gene therapy of SCID-X1 patients. J Clin Invest. 118, 3143-3150 (2008).
- Cartier, N., et al. Hematopoietic stem cell gene therapy with a lentiviral vector in X-linked adrenoleukodystrophy. Science. 326, 818-823 (2009).
- Matrai, J., et al. Hepatocyte-targeted expression by integrase-defective lentiviral vectors induces antigen-specific tolerance in mice with low genotoxic risk. Hepatology. 53, 1696-1707 (2011).
- Penaud-Budloo, M., et al. Adeno-associated virus vector genomes persist as episomal chromatin in primate muscle. Journal of virology. 82, 7875-7885 (2008).
- Kaeppel, C., et al. A largely random AAV integration profile after LPLD gene therapy. Nature medicine. 19, 889-891 (2013).
- Voigt, K., et al. Retargeting sleeping beauty transposon insertions by engineered zinc finger DNA-binding domains. Mol Ther. 20, 1852-1862 (2012).
- Yanez-Munoz, R. J., et al. Effective gene therapy with nonintegrating lentiviral vectors. Nature medicine. 12, 348-353 (2006).
- Boztug, K., et al. Stem-Cell Gene Therapy for the Wiskott-Aldrich Syndrome. N Engl J Med. 363, 1918-1927 (2010).
- Deichmann, A., et al. Vector integration is nonrandom and clustered and influences the fate of lymphopoiesis in SCID-X1 gene therapy. J Clin Invest. 117, 2225-2232 (2007).
- Schwarzwaelder, K., et al. Gammaretrovirus-mediated correction of SCID-X1 is associated with skewed vector integration site distribution in vivo. J Clin Invest. 117, 2241-2249 (2007).
- Gabriel, R., et al. An unbiased genome-wide analysis of zinc-finger nuclease specificity. Nat Biotechnol. 29, 816-823 (2011).
- Dieter, S. M., et al. Distinct types of tumor-initiating cells form human colon cancer tumors and metastases. Cell Stem Cell. 9, 357-365 (2011).
- Montini, E., et al. Hematopoietic stem cell gene transfer in a tumor-prone mouse model uncovers low genotoxicity of lentiviral vector integration. Nature biotechnology. 24, 687-696 (2006).
- Zavidij, O., et al. Stable long-term blood formation by stem cells in murine steady-state hematopoiesis. Stem Cells. 30, 1961-1970 (2012).
- Martins, V. C., et al. Thymus-autonomous T cell development in the absence of progenitor import. J Exp Med. 209, 1409-1417 (2012).
- Arens, A., et al. Bioinformatic clonality analysis of next-generation sequencing-derived viral vector integration sites. Hum Gene Ther Methods. 23, 111-118 (2012).

