A subscription to JoVE is required to view this content. Sign in or start your free trial.
JoVE Journal
Biology
An Efficient Method for Quantitative, Single-cell Analysis of Chromatin Modification and Nuclear Architecture in Whole-mount Ovules in Arabidopsis
Chapters
- 00:05Title
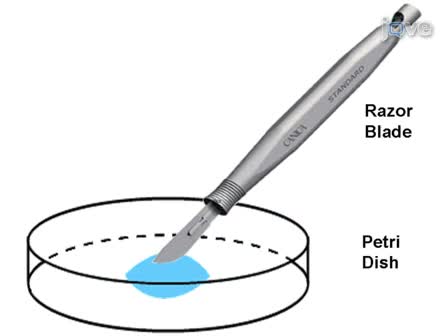
- 01:15Tissue Fixation, Dissection and Embedding
- 03:21Tissue Processing
- 05:41Immunostaining and Imaging
- 07:32High-Resolution 3D Quantitative Imaging in Ovules
- 08:40Conclusion
We provide here an efficient and reliable protocol for immunostaining, Fluorescence in situ Hybridization, DNA staining followed by quantitative, high-resolution imaging in whole-mount Arabidopsis thaliana ovules. This method was successfully used to analyze chromatin modifications and nuclear architecture.