Evaluation of Stem Cell Properties in Human Ovarian Carcinoma Cells Using Multi and Single Cell-based Spheres Assays
Summary
In vitro spheres assays are commonly used to identify cancer stem cells. Here we compare single with multi cell-based spheres assays. The more laborious single cell-based assays or methylcellulose supplementation give more accurate results while multi cell-based assays performed in liquid medium can be highly influenced by cell density.
Abstract
Years of research indicates that ovarian cancers harbor a heterogeneous mixture of cells including a subpopulation of so-called “cancer stem cells” (CSCs) responsible for tumor initiation, maintenance and relapse following conventional chemotherapies. Identification of ovarian CSCs is therefore an important goal. A commonly used method to assess CSC potential in vitro is the spheres assay in which cells are plated under non-adherent culture conditions in serum-free medium supplemented with growth factors and sphere formation is scored after a few days. Here, we review currently available protocols for human ovarian cancer spheres assays and perform a side-by-side analysis between commonly used multi cell-based assays and a more accurate system based on single cell plating. Our results indicate that both multi cell-based as well as single cell-based spheres assays can be used to investigate sphere formation in vitro. The more laborious and expensive single cell-based assays are more suitable for functional assessment of individual cells and lead to overall more accurate results while multi cell-based assays can be strongly influenced by the density of plated cells and require titration experiments upfront. Methylcellulose supplementation to multi cell-based assays can be effectively used to reduce mechanical artifacts.
Introduction
There is increasing evidence that ovarian carcinomas are comprised of heterogeneous mixtures of cells and harbor so-called “cancer stem cells” (CSCs) responsible for disease initiation, maintenance and relapse after conventional cytotoxic therapies1-3. Therefore, the development of molecular strategies targeting ovarian CSCs is an important goal and promises to improve the therapy of ovarian cancer patients.
A pre-requisite for the understanding of the molecular features of CSCs is their reliable isolation from the non-CSCs. However, identification of ovarian CSCs appears challenging. While CD133 expression and aldehyde dehydrogenase (ALDH) activity4,5 have been reported to mark ovarian CSCs, some data indicate that these markers are unstable6. Consistently, in ovarian cancer, other than for example in breast carcinoma7, expression of ALDH1 associates with favorable outcome8 and expression of the proposed stem cell marker CD44 variant has no prognostic value9. More recently, we have shown that expression of the embryonic stem cell protein SOX2 confers stemness to ovarian carcinoma cells10 and high SOX2 expression associates with clinically aggressive ovarian and breast carcinomas11,12. Therefore, in this report we use a lentiviral reporter construct containing a red fluorescence protein (RFP) whose expression is controlled by a SOX2 regulatory region, as a method to isolate putative ovarian CSCs.
By definition, CSCs can both self-renew and differentiate, giving rise to all tumor cell types. Putative CSC populations need to be analyzed in functional assays performed in vivo. For obvious reasons, in human cells such functional tests are confined to xenograft assays, comprising mostly transplantation of human tumor cells into immuno-compromised mice10,13.
An alternative in vitro method was offered by Brent Reynolds and Sam Weiss who firstly reported the so-called neurosphere assay as a surrogate assay evaluating stem potential in neural cells14. Dontu and colleagues later confirmed the use of this assay for evaluation of stem cell potential in breast cells15,16. Here, human mammary cells were plated in different numbers in serum-free medium supplemented with epidermal growth factor (EGF), basic fibroblast growth factor (bFGF), B-27 and heparin and cultured under non-adherent conditions for seven to ten days before sphere formation was scored by microscopy. Following this protocol with some adjustments in cell numbers, growth medium and supplements, several groups have explored in vitro stem cell potential from several cancer types such as breast17, brain18, pancreas19 and colon20 tumors. In ovarian carcinoma, we have recently reported feasibility of the spheres assay and compared its results to those collected in in vivo murine xenograft models10. We found that overexpression of the stem cell protein SOX2 enhanced both in vitro sphere formation as well as in vivo tumorigenicity of human ovarian carcinoma cells10. However, the frequency of sphere-initiating cells was higher than the frequency of tumor-initiating cells measured in vivo10 suggesting that either the sphere assay may lead to false positive results due to technical reasons or, alternatively, the in vivo assay may be inefficient and result in false negative results.
In this report, we analyze multi cell-based ovarian spheres assays in more detail, review the different protocols available in the literature and compare them to a single cell-based assay. We show that the single cell-based assay provides more accurate and reproducible results than multi cell-based assays, which can be highly influenced by the density of plated cells unless methylcellulose is added to the cultures to immobilize cells. However, also in single cell-based assays, in vitro sphere-initiating potential is observed at higher frequency than in vivo tumor-initiating potential.
Protocol
1. Generation of OVCAR-3 Human ovarian Carcinoma Cells Stably Transduced with Lentiviruses Containing the SOX2 Regulatory Region Reporter Construct
- Generate lentiviral particles by transfecting the HEK 293T-packaging cell line with a reporter construct recognizing a SOX2 regulatory region as described10,21.
NOTE: The reporter construct further contains a destabilization domain of the ProteoTuner Shield System ahead of the tdTomato fluorescence protein. Shield1 binds to the destabilization domain thereby preventing the proteasome to degrade the fluorescence protein22. - Transduce OVCAR-3 cells with lentiviral particles over a time period of 24 hr. Afterwards, remove the viral supernatant and wash the cells with phosphate buffered saline (PBS) and cultured in complete medium (RPMI supplemented with 10% FBS, 100 U/ml penicillin, 100 μg/ml streptomycin).
- 48 hr later, 10 μg/ml puromycin were added to the cultures and maintained for 5 days to allow selection of properly transduced cells.
2. Preparation of Cell Sorting and Plating
- Add Shield1 at 1:1,000 dilution 24 hr prior to cell sorting. Use stably transduced OVCAR-3 cells without Shield1 treatment as negative controls (Figure 1). Aspirate media from flask, wash cells with 1x PBS and trypsinize cells with 0.05% Trypsin-EDTA for 3 min.
- Stop trypsin by using complete medium (see above), count cell numbers, centrifuge cells at 300 x g at RT (15 – 25 °C) for 5 min.
- Decant supernatant and resuspend cells carefully in 0.5 – 1 ml sterile PBS.
- Use 40 µm cell strainer cap filter to obtain single-cell suspension.
- Adjust cell count to 5 million cells per ml.
- Prepare ultra low-attachment 96-well plates with 100 μl spheres medium (MEGM supplemented with growth factors, cytokines, and supplements, B-27, heparine-sodium; or DMEM/F12 supplemented with growth factors, cytokines, and supplements, B-27, heparine-sodium with or without addition of 1% methylcellulose, see also Table 1). Optionally add antibiotics to the medium at a concentration of 100 U/ml penicillin and 100 μg/ml streptomycin to minimize the risk of possible contamination.
- Sort RFP+ and RFP- cells into prepared 96-well plates from above, 1 cell per well (single cell-based spheres assay) and 100 cells per well (multi cell-based spheres assay), respectively. Perform sort on commercially available cell sorter (see Materials) using single cell mode, Sort setup: 100 μ nozzle, sheath pressure 20 psi, and yield mask 0, purity mask 32, phase mask 16.
- Assess plating efficiency by microscopically scoring wells containing cells (for the single cell-based assay) and by counting cell numbers in individual wells (for the multi cell-based assay; Figure 2).
- Incubate cells under standard conditions in spheres medium (for composition see step 2.6) at 37 °C and 5% CO2. Supplement daily bFGF (20 ng/ml) and EGF (20 ng/ml).
- After one week, count numbers of emerging tumor spheres using a standard microscope with 4X or 10X magnification and a fluorescence microscope to detect fluorescence signal from the integrated reporter system. Count spheres with a diameter exceeding 100 μm as “large” spheres, and spheres with a diameter 50 – 100 μm as “small” spheres (Figure 3). Be sure that you count real spheres and not cell clusters.
NOTE: In single cell-based assays sphere formation is easier to score microscopically after 10 (versus 7) days of culture. - Calculate the proportion of sphere-forming cells in RFP+ and respectively RFP- cells in single cell-based assays (one 96-well plate for each individual experiment) or multi cell-based spheres assays (one well for each individual experiment) as presented by Shaw et al.16
NOTE: Proportion of sphere forming cells (%) = (number of spheres) / (number of seeded cells) x 100
3. Serial Passaging of Spheres
- Place the content of each well in an appropriate sterile tube and centrifuge at 300 x g for 10 min at RT. For multi cell-based spheres assays, collect together the spheres from one well. Wash the well 3 – 5 times with PBS and centrifuge 2 min longer. For single cell-based spheres assays, collect individual spheres. Due to the low numbers of cells, use 1.5 ml tubes for centrifugation and washing steps.
- Remove supernatant and resuspend pellet in 200 µl of 0.05% Trypsin-EDTA.
- In order to achieve optimal cell separation, incubate the cell suspension at 37 °C for 5 min in a soft shaker. Optimize trypsinization time for your cell line to lower cell death rate: if no large spheres is visible triturate gently using a 100 μl pipette tip. In case large spheres are still present, incubate with trypsin another 3 min and then proceed to the trituration step. In case of single cell-based assays make sure to optimize the time for optimal cell yield of living cells during the trypsinization step.
- To inactivate the trypsin, add 500 µl complete medium and centrifuge at 300 x g for 10 min, in the case of single cell assays, centrifuge for additional 2 min.
- Remove supernatant and resuspend cells carefully in spheres medium.
- Use a 40 µm cell strainer cap filter to obtain a single-cell suspension.
- In the case of using RFP+ and RFP- cells in multi cell-bassed assays, assess the percentage of fluorescent cells in each well after resuspension via flow cytometer analysis.
- For serial replating assays of single cells, seed 1 cell per well into a new ultra low-attachment 96-well plate prepared as detailed above. From one individual sphere, seed approximately 20 individual wells. For replating assays of multi cell-based primary spheres, seed cells obtained from one well of primary spheres into a new 96-well plate and count the cell numbers next day by microscopy.
- Assess the proportion of sphere-forming cells in secondary spheres assays using the formula described in step 2.11.
4. Result Analysis
- Analyze results from experiments performed in independent triplicates and use two-sided Student’s t-Test to analyze normally distributed values and otherwise Mann-Whitney-Tests for statistical analysis.
Representative Results
In conventional spheres assays, nearly 40% of RFP+ OVCAR-3 cells vs. 20% of RFP- cells gave rise to an individual tumor sphere in the primary spheres assay (Figure 4A). Moreover, spheres formed by RFP+ cells were larger in size than those formed by RFP- cells.
When plated in single cell-based assays, RFP+ cells also formed more spheres than RFP- cells, confirming the results above. However, there was a tendency towards generation of fewer spheres per plated in the single versus the multi cell-based assay (Figure 4A,B), indicating that in this assay sphere formation may be biased through technical artifacts such as mechanical sphere fusion or dissociation in the non-adherent culture medium.
To further explore these aspects, we compared the influence of cell plating density on spheres formation. We plated cells using limiting dilution from 1,000 cells to 1 cell per well in 96-well plates and found that the numbers of emerging spheres were highly dependent on the numbers of initially plated cells. Surprisingly, higher numbers of spheres were counted from lower numbers of plated cells in both MGEM and DMEM/F12-based media, demonstrating that indeed plating modalities highly bias results in this assay (Figure 5). In contrast, when cells were immobilized by adding 1% methylcellulose to DMEM/F12-based spheres medium23,24 the efficiency of sphere formation was mostly independent of cell density.
To explore the influence of spheres culture conditions on CSC properties, we analyzed the percentage of cells expressing red fluorescence signal after 7 days of incubation in the spheres assay. We found that in multi cell-based spheres cultures approximately 35% of the cells from RFP+ spheres lost their red fluorescence signal after seven days of culture (Figure 6A), suggesting that they have undergone differentiation, while 65% retained a RFP+ signal suggesting self-renewal capacity. In contrast, cells from spheres generated from initially RFP- cells remained RFP negative (Figure 6A), indicating that they cannot re-establish stem cell potential under these conditions.
Fluorescence microscopy performed on spheres generated from single cells confirmed these results showing that single spheres derived RFP+ cells contained both RFP+ and RFP- cells while spheres derived from RFP- cells remained negative for the red signal.
Similar results were observed in replating assays from both conditions.
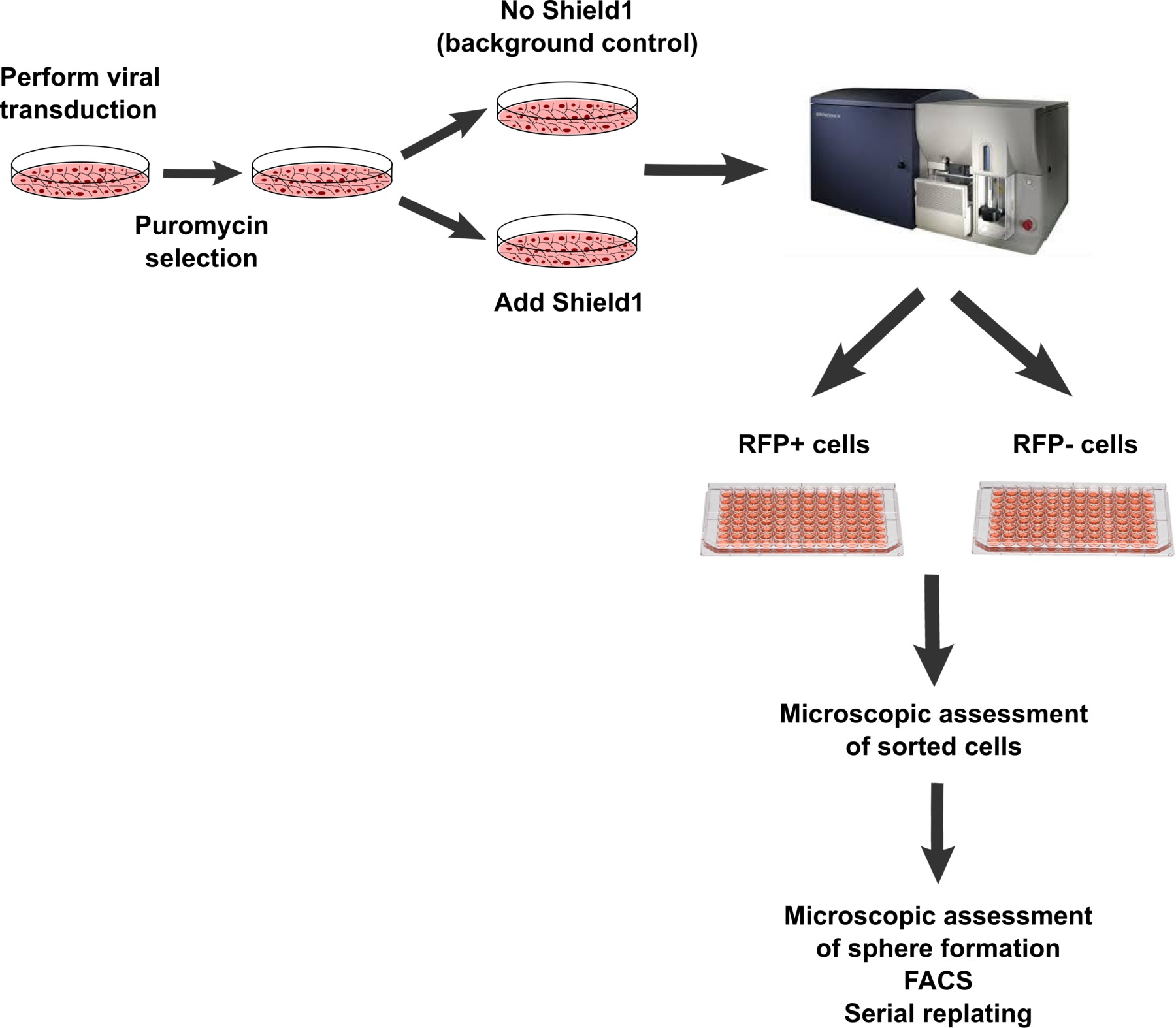
Figure 1. Workflow of lentiviral transduction, selection and sorting of RFP+ and RFP- cells. After lentiviral transduction and positive selection of successfully transduced cells via puromycin exposure, RFP- and RPF+ cells are sorted by FACS into individual wells of a 96-well plate in spheres medium. For multi cell-based spheres assays, 100 cells are placed into one well. Plating efficiency is assessed by microscopy performed after sorting. Spheres were scored by microscopy after seven to ten days, dissociated into single cells, analyzed via flow cytometry and replated into secondary spheres. Please click here to view a larger version of this figure.
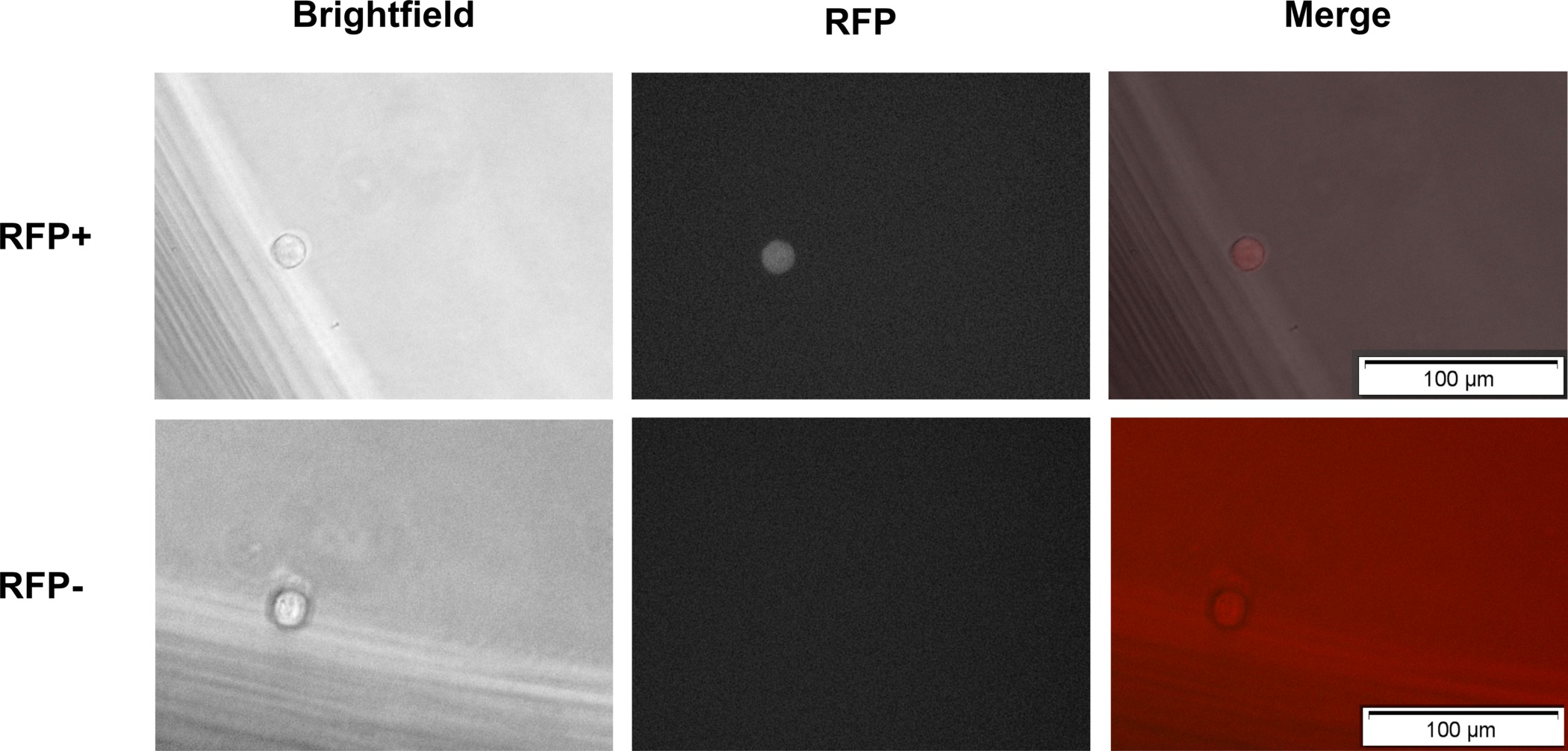
Figure 2. Imaging of sorted RFP+ and RFP- cells after plating. Single RFP+ and RFP- cells sorted into each well of a 96-well plate are analyzed for correct plating by using a (fluorescent) microscope. Please click here to view a larger version of this figure.
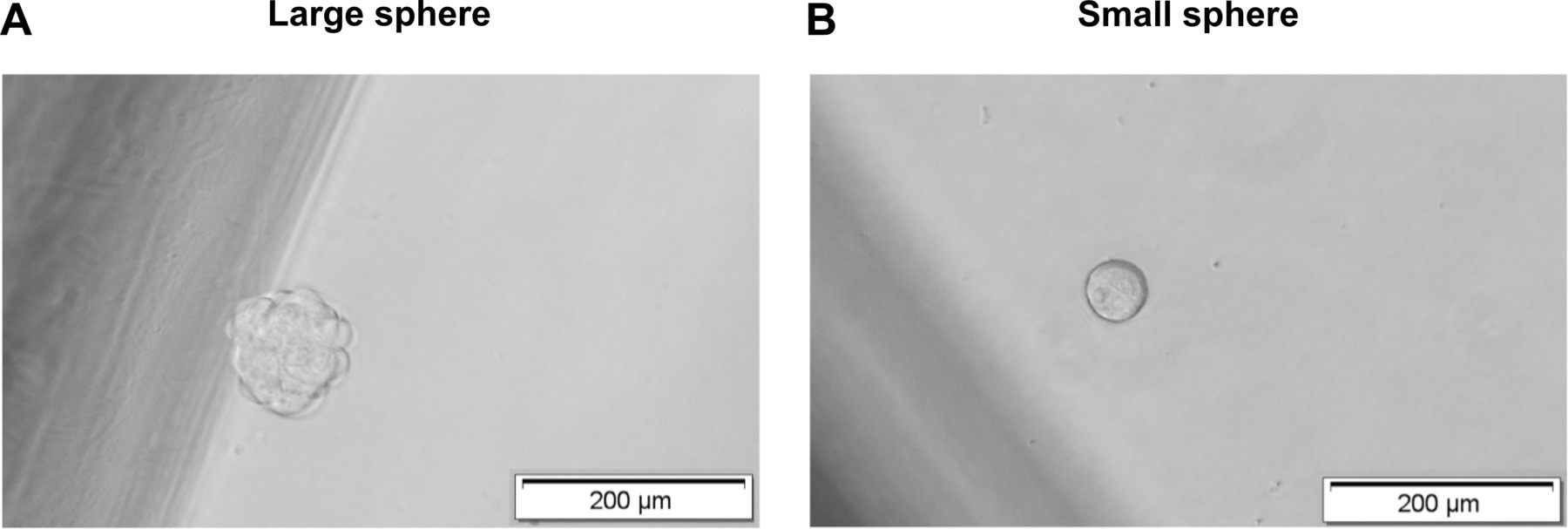

Figure 3. Analysis of spheres formation in single cell-based assays. Spheres formation is analyzed after seven to ten days. (A) Large (diameter > 100 µm) and (B) small (diameter 50 – 100 µm) spheres are distinguished microscopically based on size. Please click here to view a larger version of this figure.
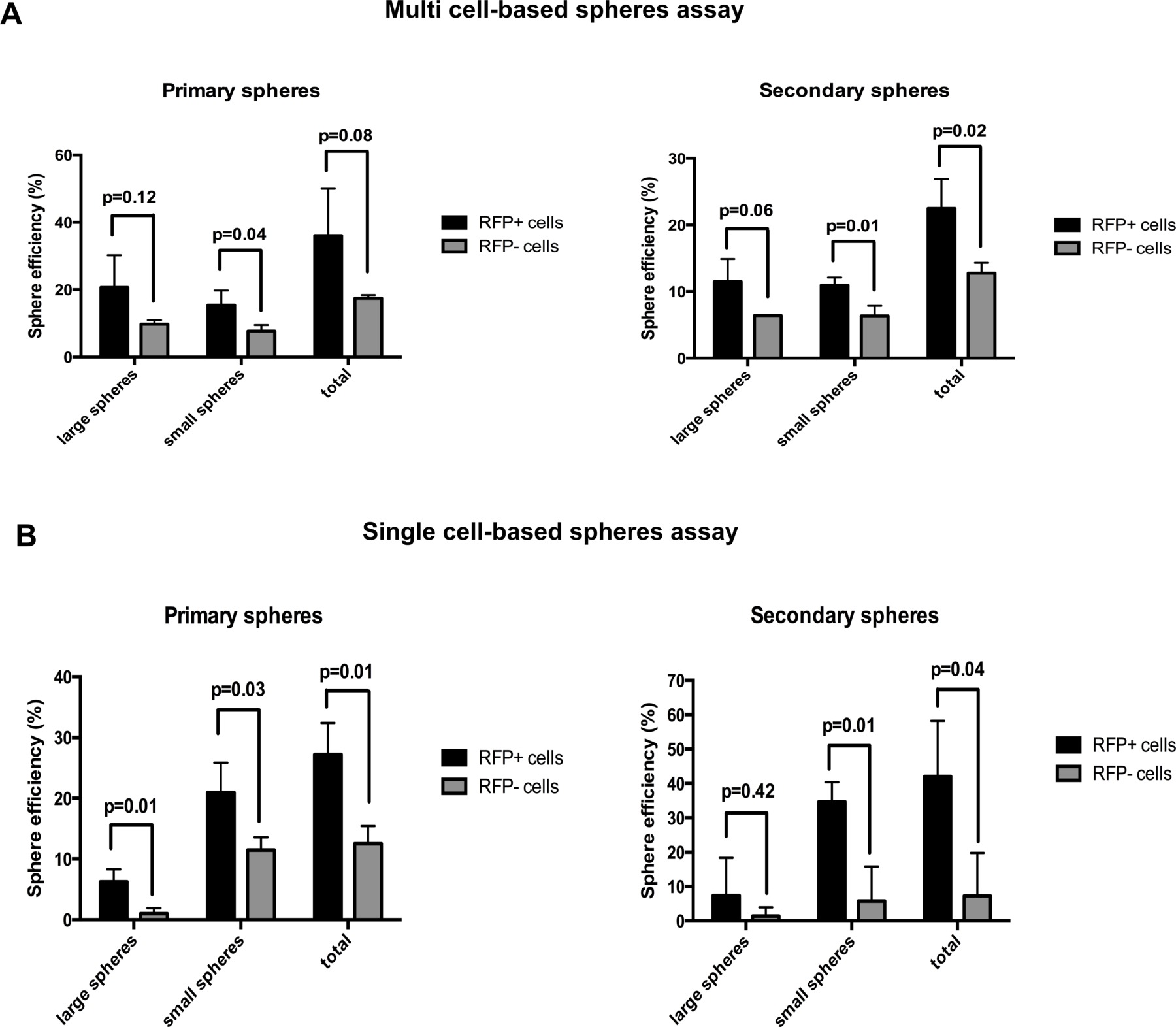
Figure 4. Efficiency of tumor spheres formation from RFP+ and RFP- OVCAR-3 cells in multi versus single-cell based spheres assays. Comparison of primary and secondary sphere efficiency from OVCAR-3 cells as assayed in multi (A) versus single cell-based spheres assays (B). Please click here to view a larger version of this figure.

Figure 5. Cell plating density strongly impacts sphere counts from OVCAR-3 cells in the multi cell-based spheres assay performed in liquid but not in methylcellulose supplemented cultures. Use of different cell densities and growth media have been reported in the literature for ovarian cancer spheres assays. To analyze possible biases introduced by these variables, cells are plated at different densities in 200 µl of different spheres culture media (MGEM, DMEM/F12 with all supplements as detailed in the protocol section, or DMEM/F12 with all supplements and containing 1% methylcellulose) and sphere formation is scored after 7 days (A). Shown in (B) are microscopy pictures of cells plated at different densities taken one day after plating in DMEM/F12 spheres culture medium without methylcellulose. Note the cell clusters emerging at high cellular density as opposed to single cells seen in low density plates. Scale bar for pictures: 50 µm. Please click here to view a larger version of this figure.
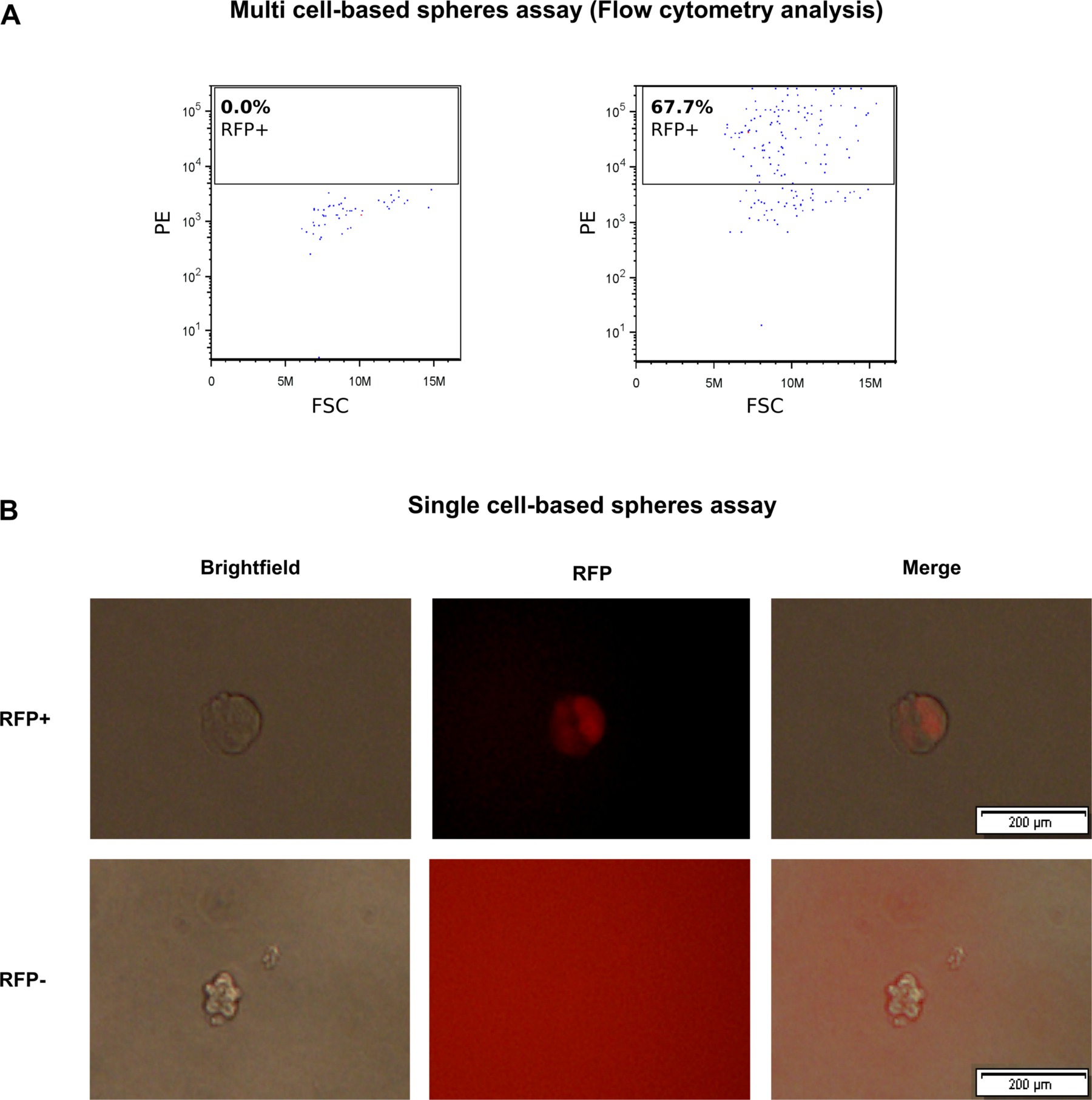
Figure 6. Analysis of RFP signal in tumor spheres formed from RFP+ and respectively RFP- OVCAR-3 cells. (A) Flow cytometry analysis for RFP signal in dissociated spheres derived from RFP+ and RFP- cells (multi cell-based spheres assay); (B) Microscopy of spheres derived from RFP+ and RFP- cells (single cell-based spheres assay) reveals heterogeneous RFP signal in spheres derived from RFP+ but not from RFP- cells. Pictures were taken at day 7 for conventional spheres and day 10 for single cell-based sphere assays. Note the larger size of spheres derived from RFP+ putative CSCs. Please click here to view a larger version of this figure.
| Human ovarian cancer cell source | Basic medium | Supplements | Authors |
| OVCAR-3, Caov-3, primary material | MEGM | 20 ng/ml rEGF, 20 ng/ml bFGF, B-27, 4 μg/ml heparin, hydrocortisone, insulin (SingleQuot kit) | Bareiss et al. |
| SKOV3 | DMEM/F12 | 5 µg/ml insulin, 10 ng/ml rEGF, 10 ng/ml bFGF, 12 ng/ml LIF, 0.3% BSA | Li Ma et al. |
| A2780 | DMEM/F12 | 5 µg/ml insulin, 20 ng/ml rEGF, 2% B-27, 0.4% BSA | Haiwei Wang et al. |
| SKOV3 | DMEM/F12 | 5 µg/ml insulin, 20 ng/ml rEGF, 10 ng/ml bFGF, 2% B-27, 1 ng/ml hydrocortisone | Yong-Rui Du et al. |
| A2780, primary material | DMEM/F12 | 5 µg/ml insulin, 20 ng/ml rEGF, 10 ng/ml bFGF, 0.4% BSA | T. Xiang et al. |
| Primary material | DMEM/F12 | 5 µg/ml insulin, 10 ng/ml rEGF, 10 ng/ml bFGF, 12 ng/ml LIF, 0.3% BSA | Te Liu et al. |
| MLS | DMEM/F12 | 10 ng/ml insulin, 20 ng/ml rEGF, 20 ng/ml bFGF, 2% B-27 | Soritau et al. |
| 3AO | DMEM/F12 | 1 mg/ml insulin, 20 ng/ml rEGF, 20 ng/ml bFGF, 2% B-27 | M. F. Shi et al. |
| Primary material | DMEM/F12 | 5 µg/ml insulin, 20 ng/ml rEGF, 10 ng/ml bFGF, 0.4% BSA | Shu Zhang et al. |
| Primary material | EBM-2 or X-VIVO | 5 µg/ml insulin, 20 ng/ml rEGF | Ilona Kryczek et al. |
| OVCAR-3 | MEGM | 20 ng/ml rEGF, 20 ng/ml bFGF, B-27, 4 μg/mL heparin | Dongming Liang et al. |
Table 1. Examples of different cell sources (cell lines and primary patient-derived tissue), media and supplements used for ovarian spheres assays in previous reports.
Discussion
Spheres cultures are a widely used method to assay cancer stem cell potential and enrich for stem-like cells in a wide range of human tumor cells15,25,26. Under these culture conditions, cancer cells that lack self-renewal ability are expected to differentiate and eventually undergo cell death. Although they may initially form cell clusters or even tumor spheres especially in primary assays, they are not able to sustain sphere-forming ability upon serial replating due to lack of self-renewing properties. Spheres assays are used as surrogate assays to identify CSCs and evaluate their frequency in whole tumor cell populations.
However, substantial variability can be observed between spheres assays performed following different published protocols5,10,27-35 (Table 1). In our laboratory, we have previously published sphere formation from human ovarian carcinoma cells using MEGM supplemented with B-27, bFGF, Heparin and SingleQuotTM (containing insulin, rEGF and hydrocortisone). Other labs use whole DMEM/F12 Medium, while some add only B-27 and rEGF. In this report, multi cell-based spheres assay in OVCAR-3 cells were therefore performed using different conditions. Using MEGM or DMEM/F12 with all supplements no significant difference in sphere formation was observed in these cells (Figure 5A). In addition, some labs have speculated that EGF and FGF may be quickly degraded and have established protocols adding these growth factors daily to the medium. We therefore compared spheres assays performed in a medium in which EGF and FGF was added only at the beginning of the spheres assays with daily addition of EGF and FGF to the cell culture, and we find these assays to yield equivalent results in OVCAR-3 cells (data not shown), suggesting that the expensive and laborious daily supplementation with EGF and FGF may not always be necessary. Whether these results are applicable to cells from other ovarian cancer cell lines or primary samples, or under different experimental conditions remains to be determined.
However, we observed a substantial bias in the numbers of scored spheres introduced by another tested variable, the cell plating density. Surprisingly, wells seeded with lower numbers of cells showed higher numbers of spheres. Limiting cell mobility by 1% methylcellulose resulted in the same efficiency of sphere formation, independent on the number of initially plated cells. These results suggest that cell clumping and sphere fusion or disaggregation can occur modifying sphere numbers and leading to inaccurate results in multi cell-based spheres assays. When cells are plated at proper density, multi cell-based assays however lead to results rather comparable to data collected in single cell-based sphere assays (Figure 4). To further explore these results we compared single and multi cell-based assays using ovarian carcinoma cells sorted into putative CSCs via a recently published lentiviral RFP expressing reporter system for a SOX2 regulatory region10. Indeed, both assays confirmed the enhanced primary and secondary sphere forming capacity of RFP+ versus RFP- cells (Figure 4A,B). Importantly, the higher numbers of spheres observed from RFP+ cells were not due to higher proliferative capacity of the SOX2 expressing cells (data not shown), which is in line with previous results showing that induction of SOX2 promotes spheres formation and in vivo tumorigenicity without accelerating cell cycle progression10.
Taken together, single cell-based spheres assays are more laborious and expensive but they result in more accurate data, which also is confirmed by the higher reproducibility of results between experiments. Since plating density highly influences results in multi cell-based suspension spheres assays, upfront titration of adequate plating density is required for each individual tumor cell type before assaying sphere formation using these assays. Alternatively, the more accurate single cell-based assays can be used upfront, or methylcellulose supplementation to improve accuracy of results by reducing mechanical artifacts. If sphere formation is compared between conditions where the genetic modification or drug treatment may severely alter viability of the cells, thereby decreasing cell density, single cell-based sphere assays may be mandatory to avoid false positive results.
In summary, under proper experimental conditions both the multi cell-based spheres assay and the single cell-based spheres assay are able to indicate differences in sphere potential between different cell populations (stem and non-stem cells). However, multi cell-based spheres assays which commonly are performed in liquid cultures are more susceptible to errors introduced by experimental design through plating density. Supplementation of methylcellulose (1%) to multi cell-based assays can limit artifacts related to cell clumping and sphere fusion. Based on these data, we recommend single cell-based spheres assays to be performed unless detailed titration analyses have been performed upfront and negative impact of experimental conditions on cell viability and thereby cell density has been ruled out. However, single cell-based spheres assays are more laborious and more expensive, and might not be required in each experimental setting. Methylcellulose-supplemented multi cell-based spheres assays may represent another alternative in some experimental settings.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This study was supported by a grant from the Baden-Württemberg Stiftung (Adult Stem Cells Program II) awarded to C.L. We thank Dr. Martina Konantz for critical input and review of the manuscript. We thank Emmanuel Traunecker and Toni Krebs from the DBM FACS Facility (University Hospital Basel) for assistance with FACS sorting.
Materials
| Name of Material/ Equipment | Company | Catalog Number | Comments/Description |
| low-Attachment-plate | Corning | 3474 | |
| MEGM | Lonza | CC-3151 | |
| Insulin | Lonza | CC-4136 | SingleQuots™ Kit |
| Hydrocortison | Lonza | CC-4136 | SingleQuots™ Kit |
| EGF | Lonza | CC-4136 | SingleQuots™ Kit |
| EGF | Sigma | E9644 | end concentration: 20 ng/ml |
| FGF | PeproTech | 100-18B | end concentration: 20 ng/ml |
| B-27 | Invitrogen/ Gibco | 17504-044 | end concentration: 1x |
| Heparin-Natrium-25000 IE | Ratiopharm | N68542.02 | dilution 1:1000 |
| Pen/Strep | Gibco | 15140-122 | |
| FCS | Gibco | 10500-064 | |
| RPMI 1640 | Gibco | 21875-034 | |
| Trypsin-EDTA | Gibco | 25300-054 | |
| Dulbecco’s PBS (1x) | Gibco | 14190-094 | |
| Shield1 | Clontech | 632189 | dilution 1:1000 |
| DMEM/F12 | Gibco | 21041-025 | |
| DMEM/F12 (powder) | Gibco | 42400-010 | |
| Methyl cellulose | Sigma | M0387 | |
| Puromycin dihydrochloride | applichem | A2856 | |
| cell sorter | BD | Aria III cell sorter | |
| FACS analyser | BD | accuri c6 flow cytometer | |
| microscope | Olympus | IX50 Osiris |
References
- Pardal, R., Clarke, M. F., Morrison, S. J. Applying the principles of stem-cell biology to cancer. Nat Rev Cancer. 3 (12), 895-902 (2003).
- Reya, T., Morrison, S. J., Clarke, M. F., Weissman, I. L. Stem cells, cancer, and cancer stem cells. Nature. 414 (6859), 105-111 (2001).
- Ahmed, N., Abubaker, K., Findlay, J. K. Ovarian cancer stem cells: Molecular concepts and relevance as therapeutic targets. Mol Aspects Med. 39, 110-125 (2014).
- Silva, I. A., et al. Aldehyde dehydrogenase in combination with CD133 defines angiogenic ovarian cancer stem cells that portend poor patient survival. Cancer Res. 71 (11), 3991-4001 (2011).
- Kryczek, I., et al. Expression of aldehyde dehydrogenase and CD133 defines ovarian cancer stem cells. Int J Cancer. 130 (1), 29-39 (2012).
- Stewart, J. M., et al. Phenotypic heterogeneity and instability of human ovarian tumor-initiating cells. Proc Natl Acad Sci U S A. 108 (16), 6468-6473 (2011).
- Ginestier, C., et al. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell. 1 (5), 555-567 (2007).
- Chang, B., et al. ALDH1 expression correlates with favorable prognosis in ovarian cancers. Mod Pathol. 22 (6), 817-823 (2009).
- Cannistra, S. A., et al. CD44 variant expression is a common feature of epithelial ovarian cancer: lack of association with standard prognostic factors. J Clin Oncol. 13 (8), 1912-1921 (1995).
- Bareiss, P. M., et al. SOX2 expression associates with stem cell state in human ovarian carcinoma. Cancer Res. 73 (17), 5544-5555 (2013).
- Pham, D. L., et al. SOX2 expression and prognostic significance in ovarian carcinoma. Int J Gynecol Pathol. 32 (4), 358-367 (2013).
- Lengerke, C., et al. Expression of the embryonic stem cell marker SOX2 in early-stage breast carcinoma. BMC Cancer. 11, 42 (2011).
- Lapidot, T., et al. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature. 367 (6464), 645-648 (1994).
- Reynolds, B. A., Weiss, S. Generation of neurons and astrocytes from isolated cells of the adult mammalian central nervous system. Science. 255 (5052), 1707-1710 (1992).
- Dontu, G., et al. In vitro propagation and transcriptional profiling of human mammary stem/progenitor cells. Genes Dev. 17 (10), 1253-1270 (2003).
- Shaw, F. L., et al. A detailed mammosphere assay protocol for the quantification of breast stem cell activity. J Mammary Gland Biol Neoplasia. 17 (2), 111-117 (2012).
- Leis, O., et al. Sox2 expression in breast tumours and activation in breast cancer stem cells. Oncogene. 31 (11), 1354-1365 (2012).
- Higgins, D. M., et al. Brain tumor stem cell multipotency correlates with nanog expression and extent of passaging in human glioblastoma xenografts. Oncotarget. 4 (5), 792-801 (2013).
- Wang, Y. J., Bailey, J. M., Rovira, M., Leach, S. D. Sphere-forming assays for assessment of benign and malignant pancreatic stem cells. Methods Mol Biol. 980, 281-290 (2013).
- Li, Y. F., Xiao, B., Tu, S. F., Wang, Y. Y., Zhang, X. L. Cultivation and identification of colon cancer stem cell-derived spheres from the Colo205 cell line. Braz J Med Biol Res. 45 (3), 197-204 (2012).
- Wu, F., et al. Identification of two novel phenotypically distinct breast cancer cell subsets based on Sox2 transcription activity. Cell Signal. 24 (11), 1989-1998 (2012).
- Banaszynski, L. A., Chen, L. C., Maynard-Smith, L. A., Ooi, A. G., Wandless, T. J. A rapid, reversible, and tunable method to regulate protein function in living cells using synthetic small molecules. Cell. 126 (5), 995-1004 (2006).
- Kawase, Y., Yanagi, Y., Takato, T., Fujimoto, M., Okochi, H. Characterization of multipotent adult stem cells from the skin: transforming growth factor-beta (TGF-beta) facilitates cell growth. Exp Cell Res. 295 (1), 194-203 (2004).
- Walia, V., et al. Loss of breast epithelial marker hCLCA2 promotes epithelial-to-mesenchymal transition and indicates higher risk of metastasis. Oncogene. 31 (17), 2237-2246 (2012).
- Leung, E. L., et al. Non-small cell lung cancer cells expressing CD44 are enriched for stem cell-like properties. PLoS One. 5 (11), e14062 (2010).
- Bertolini, G., et al. Highly tumorigenic lung cancer CD133+ cells display stem-like features and are spared by cisplatin treatment. Proc Natl Acad Sci U S A. 106 (38), 16281-16286 (2009).
- Ma, L., Lai, D., Liu, T., Cheng, W., Guo, L. Cancer stem-like cells can be isolated with drug selection in human ovarian cancer cell line SKOV3. Acta Biochim Biophys Sin (Shanghai). 42 (9), 593-602 (2010).
- Wang, H., Zhang, Y., Du, Y. Ovarian and breast cancer spheres are similar in transcriptomic features and sensitive to fenretinide). Biomed Res Int. 2013, 510905 (2013).
- Du, Y. R., et al. Effects and mechanisms of anti-CD44 monoclonal antibody A3D8 on proliferation and apoptosis of sphere-forming cells with stemness from human ovarian cancer. Int J Gynecol Cancer. 23 (8), 1367-1375 (2013).
- Xiang, T., et al. Interleukin-17 produced by tumor microenvironment promotes self-renewal of CD133 cancer stem-like cells in ovarian cancer. Oncogene. , (2013).
- Liu, T., Cheng, W., Lai, D., Huang, Y., Guo, L. Characterization of primary ovarian cancer cells in different culture systems. Oncol Rep. 23 (5), 1277-1284 (2010).
- Soritau, O., et al. Enhanced chemoresistance and tumor sphere formation as a laboratory model for peritoneal micrometastasis in epithelial ovarian cancer. Rom J Morphol Embryol. 51 (2), 259-264 (2010).
- Shi, M. F., et al. Identification of cancer stem cell-like cells from human epithelial ovarian carcinoma cell line. Cell Mol Life Sci. 67 (22), 3915-3925 (2010).
- Zhang, S., et al. Identification and characterization of ovarian cancer-initiating cells from primary human tumors. Cancer Res. 68 (11), 4311-4320 (2008).
- Liang, D., et al. The hypoxic microenvironment upgrades stem-like properties of ovarian cancer cells. BMC Cancer. 12, 201 (2012).

